| Deletions are marked like this. | Additions are marked like this. |
| Line 3: | Line 3: |
| '''AnatomiCuts''' is an unsupervised hierarchical clustering that uses an anatomical similarity metric. |
'''AnatomiCuts''' is an unsupervised hierarchical clustering that uses an anatomical similarity metric. [[File:rainbow_brain.png|thumb|right|upright=0.35]] |
| Line 16: | Line 17: |
| == Running AnatomiCuts == |
= Running AnatomiCuts = |
| Line 18: | Line 20: |
| Line 25: | Line 26: |
| = References = |
AnatomiCuts
AnatomiCuts is an unsupervised hierarchical clustering that uses an anatomical similarity metric. [[File:rainbow_brain.png|thumb|right|upright=0.35]]
Inputs
- streamllines file (*.trk) - a segmentation image, preferable including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc.mgz or wm2009parc.mgz).
It is important that both files are in the same space. To validate this you can open the streamline file and the segmentation using both Freeview and Trackvis, since Freeview does not take into account RAS orientation, and Trackvis does not take into account the image offset.
Filtering streamlines
Sometimes deterministic tractography can generate spurious streamlines outside the brain. To exclude them we can use streamlineFilter. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines.
Running AnatomiCuts
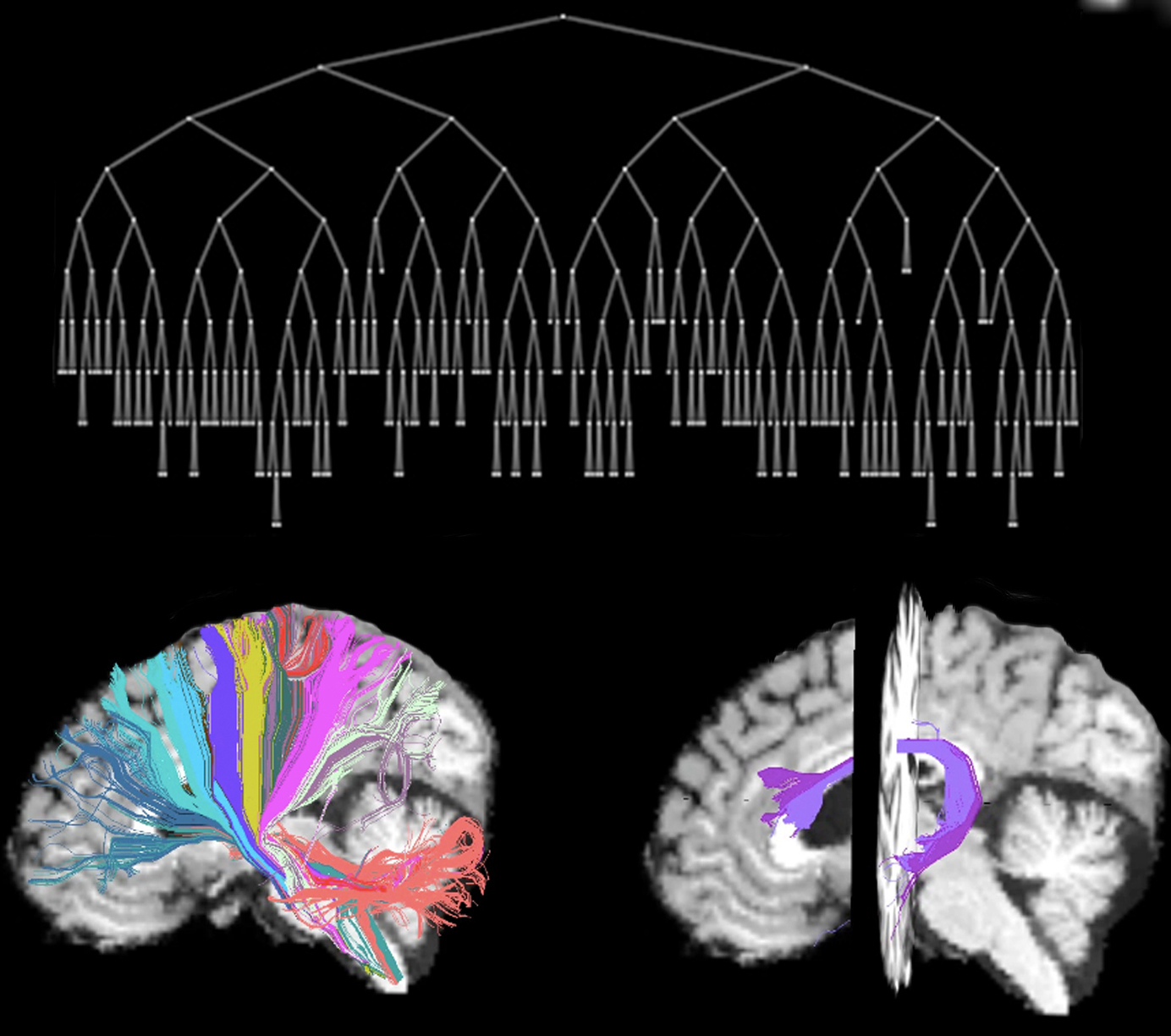
References
AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity. Siless V., Chang K., Fischl B., Yendiki A.. NeuroImage 2018.
