Bayesian Segmentation with Histological Atlas
This functionality is available on development versions newer than October 26th 2023
Author: Juan Eugenio Iglesias
E-mail: jiglesiasgonzalez [at] mgh.harvard.edu
Rather than directly contacting the author, please post your questions on this module to the FreeSurfer mailing list at freesurfer [at] nmr.mgh.harvard.edu
The manuscript describing this module is still in preparation:
Contents
- General Description
- Installation
- Usage
- Frequently asked questions (FAQ)
1. General Description
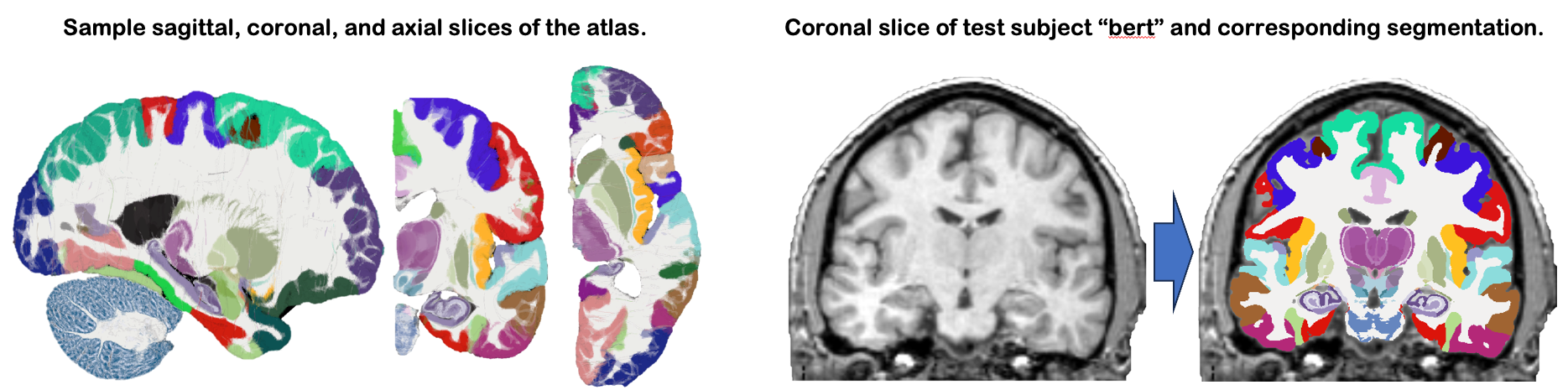
This module uses our new probabilistic atlas of the human brain to segment 333 distinct ROIs per hemisphere on in vivo scans. Segmentation relies on a Bayesian algorithm and is thus robust against changes in MRIi pulse sequence (e.g., T1-weighted, T2-weighted, FLAIR, etc). Sample slices of the atlas and the segmentation of the sample subject "bert" are shown below:
2. Installation
The first time you run this module, it will prompt you to download the atlas. Follow the instructions on the screen to obtain the files.
3. Usage
To segment a brain MRI scan,
mri_histo_atlas_segment INPUT_SCAN OUTPUT_DIRECTORY GPU THREADS
where:
INPUT_SCAN: scan to process, in mgz or nii(.gz) format.
OUTPUT_DIRECTORY: directory where segmentations, volume files, etc, will be created (more on this below).
GPU: set to 1 to use the GPU (we highly recommend using a GPU to run this module; without a GPU, running this module on a single scan can take a whole day). The GPU requirements depend on the image but are about 24GB of memory. THREADS: number of CPU threads used by the code (set to -1 to use all available threads). In the output directory, you will find: Coming soon.
4. Frequently asked questions (FAQ)
