|
Size: 668
Comment:
|
Size: 3219
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| ## Please edit system and help pages ONLY in the master wiki! ## For more information, please see MoinMoin:MoinDev/Translation. ## IMPORTANT NOTE: ## When you use this page as a template for creating your project page: ## * please remove all lines starting with two hashes (##) ## * except the acl line, please keep that, but remove one hash, so it reads #acl ... ## * fix the acl line so it has the correct page instead of the sample Project/...Group ##acl Project/AdminGroup:admin,read,write,delete,revert Project/ReadWriteGroup:read,write Project/ReadGroup:read ##master-page:Unknown-Page ##master-date:Unknown-Date #format wiki #language en sdadas |
= AnatomiCuts = '''AnatomiCuts''' is an unsupervised hierarchical clustering that uses an anatomical similarity metric to cluster together white matter streamline with similar neighboring anatomical structures. {{attachment:anatomiCuts_tree.jpg||height="360px"}}{{attachment:rainbow_brain.png||height="360px"}} <<TableOfContents>> == Inputs == - streamlines file (*.trk) - segmentation image, ideally, including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc or wm2009parc). '''It is important to ensure that the white matter streamlines and the segmentation are in the same space.''' To validate this it is recommended to visualize the streamlines file and the segmentation file using '''both''' Freeview and Trackvis, since Freeview does not take into account RAS orientation, and Trackvis does not take into account the image offset. == Filtering streamlines == Sometimes deterministic tractography can generate spurious streamlines outside the brain, or very short cut off streamlines. To exclude them we can use [[streamlineFilter| streamlineFilter]]. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines. = Running AnatomiCuts = {{{ AnatomiCuts -s segmentation.nii.gz -f streamlines.vtk -labels -o outputFolder <options> }}} Where -s segmentation file such as wm2009parc.nii.gz -f streamline file in trk format. -c number of clusters. The default is 200. -n number of points to be sampled per streamline. The default is 10. -e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice, less than 300 is not recommended. -d directional neighbors to be used, the default is "a" all with 26 directions, diagonal "d" contains 14 directions and straight "s" only 6 directions. -o output folder = Finding corresponding clusters across subjects = The Hungarian algorithm allows to find one-to-one correspondences and it's implementation for AnatomiCuts at multiple levels of the tree hierarchy can be found in [[AnatomiCuts_correspondences| AnatomiCuts_correspondences]]. = Visualizing AnatomiCuts in Freeview = To load the clusters obtained with AnatomiCuts go to "File -> Load Tract Cluster" and select the AnatomiCuts output folder. {{attachment:anatomicuts_freeview.png||height="360px"}}{{attachment:anatomicuts_freeview_tree.png||height="360px"}} {{attachment:anatomicuts.gif}} = References = AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity. Siless V., Chang K., Fischl B., Yendiki A.. NeuroImage 2018. V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.” |
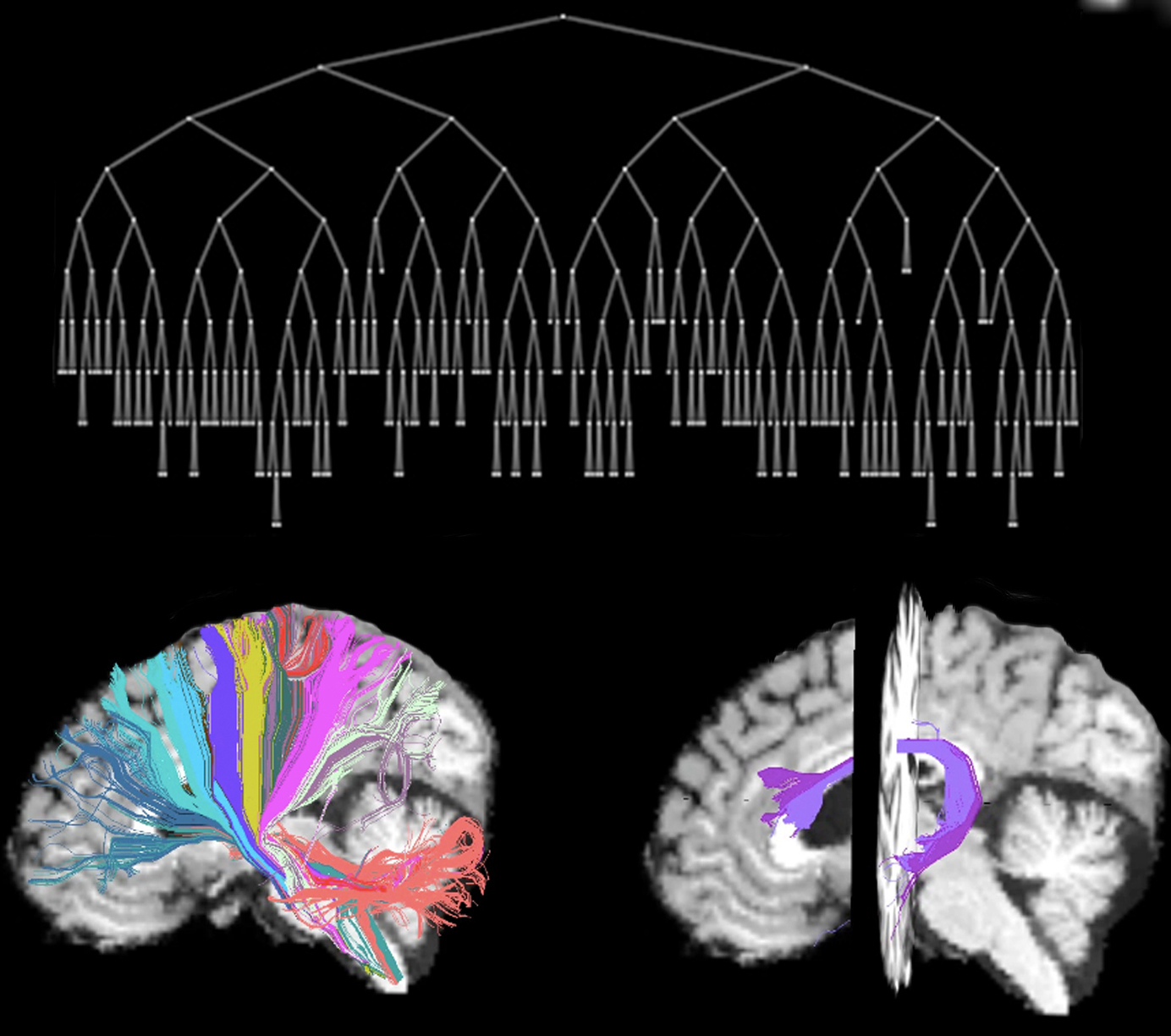
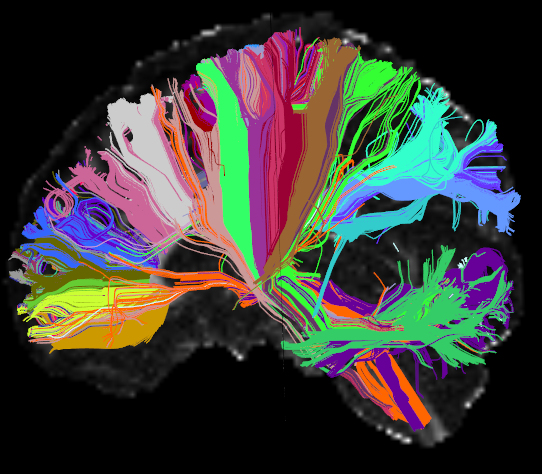
AnatomiCuts
AnatomiCuts is an unsupervised hierarchical clustering that uses an anatomical similarity metric to cluster together white matter streamline with similar neighboring anatomical structures.
Contents
Inputs
- streamlines file (*.trk)
- segmentation image, ideally, including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc or wm2009parc).
It is important to ensure that the white matter streamlines and the segmentation are in the same space. To validate this it is recommended to visualize the streamlines file and the segmentation file using both Freeview and Trackvis, since Freeview does not take into account RAS orientation, and Trackvis does not take into account the image offset.
Filtering streamlines
Sometimes deterministic tractography can generate spurious streamlines outside the brain, or very short cut off streamlines. To exclude them we can use streamlineFilter. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines.
Running AnatomiCuts
AnatomiCuts -s segmentation.nii.gz -f streamlines.vtk -labels -o outputFolder <options>
Where
-s segmentation file such as wm2009parc.nii.gz
-f streamline file in trk format.
-c number of clusters. The default is 200.
-n number of points to be sampled per streamline. The default is 10.
-e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice, less than 300 is not recommended.
-d directional neighbors to be used, the default is "a" all with 26 directions, diagonal "d" contains 14 directions and straight "s" only 6 directions.
-o output folder
Finding corresponding clusters across subjects
The Hungarian algorithm allows to find one-to-one correspondences and it's implementation for AnatomiCuts at multiple levels of the tree hierarchy can be found in AnatomiCuts_correspondences.
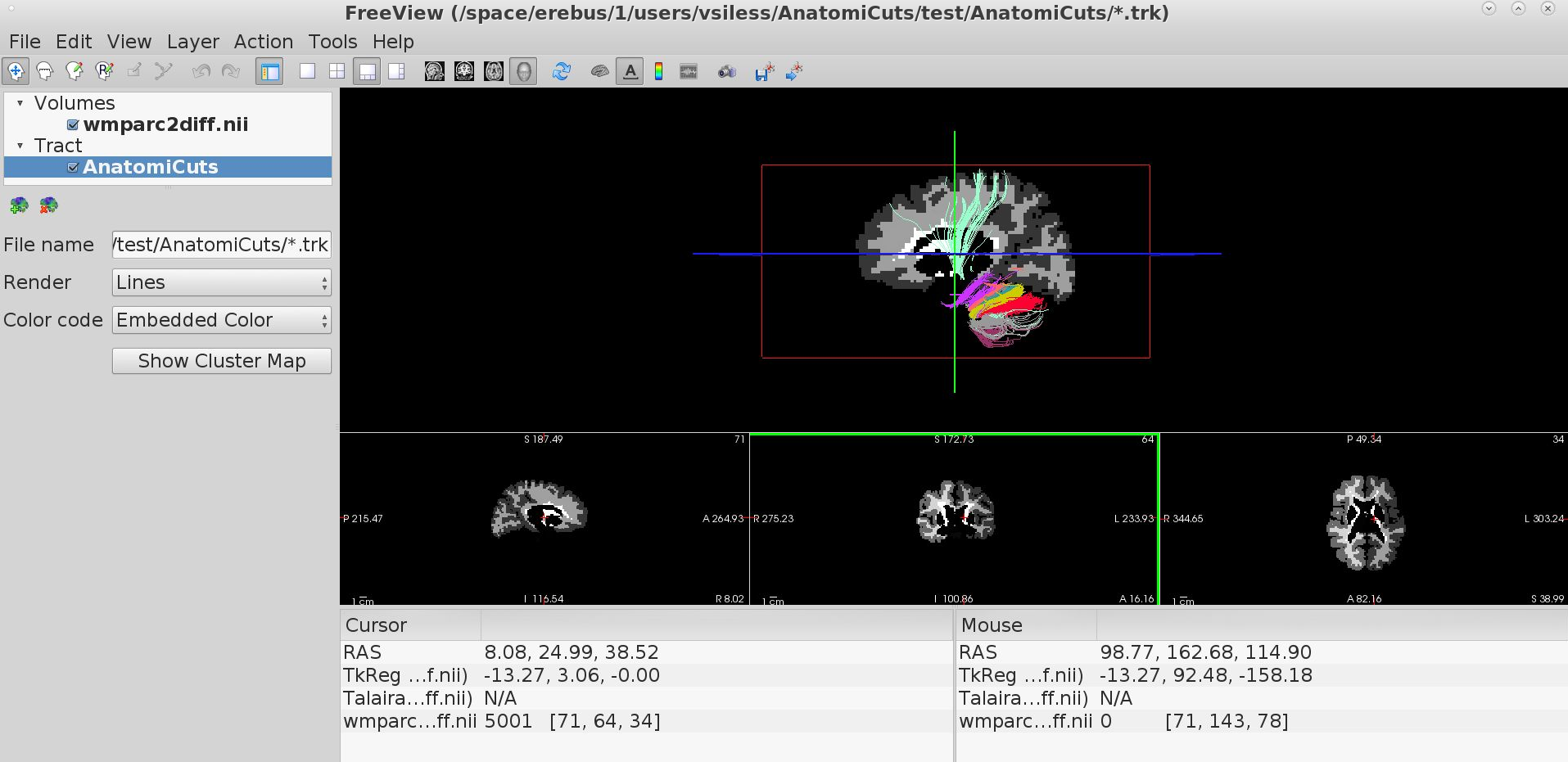
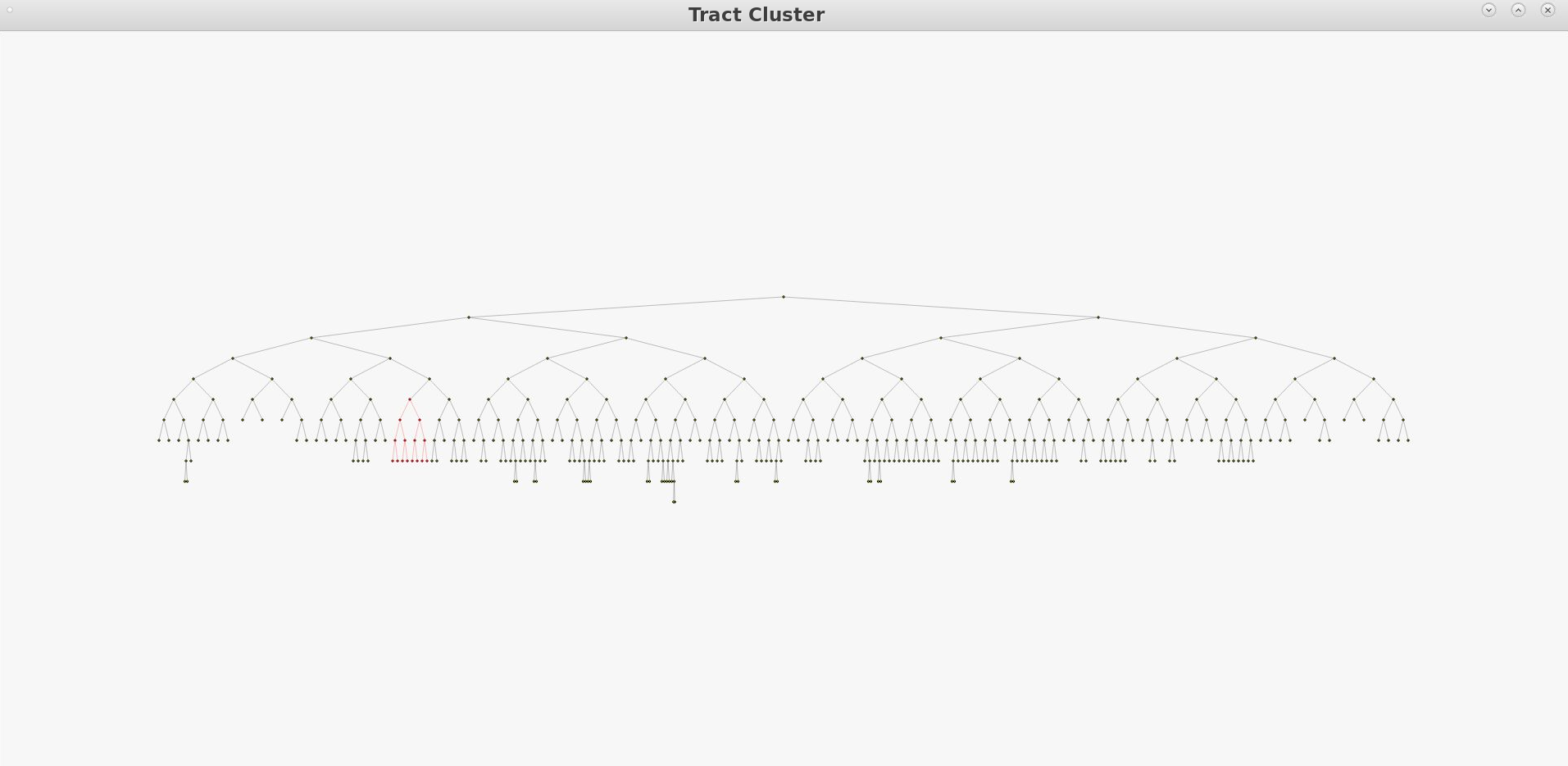
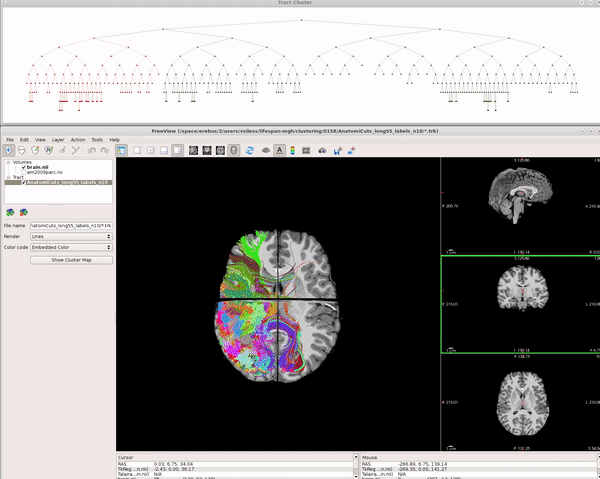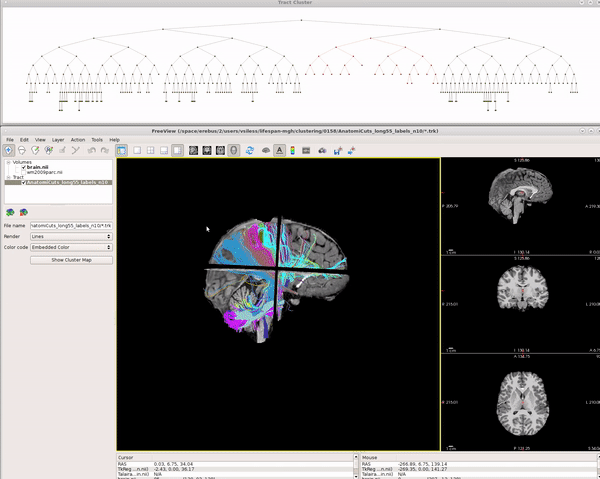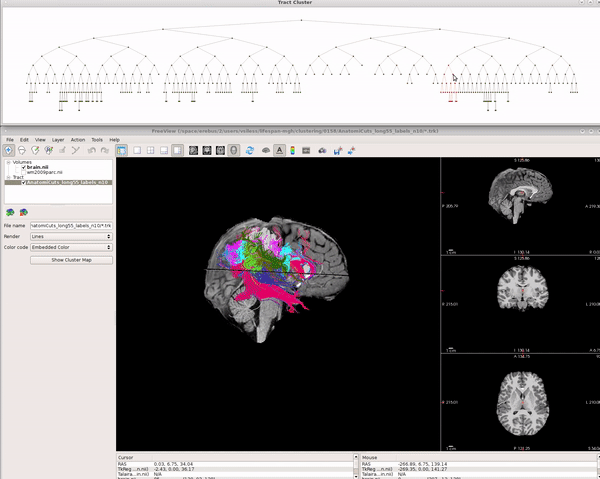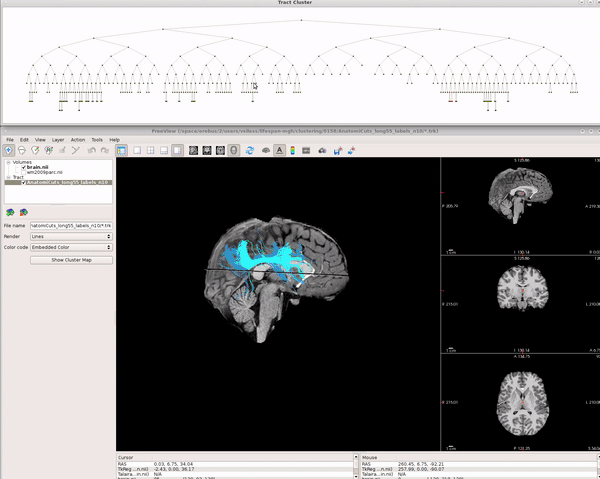
Visualizing AnatomiCuts in Freeview
To load the clusters obtained with AnatomiCuts go to "File -> Load Tract Cluster" and select the AnatomiCuts output folder.
References
AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity. Siless V., Chang K., Fischl B., Yendiki A.. NeuroImage 2018.
V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.”