|
Size: 2676
Comment:
|
Size: 4378
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
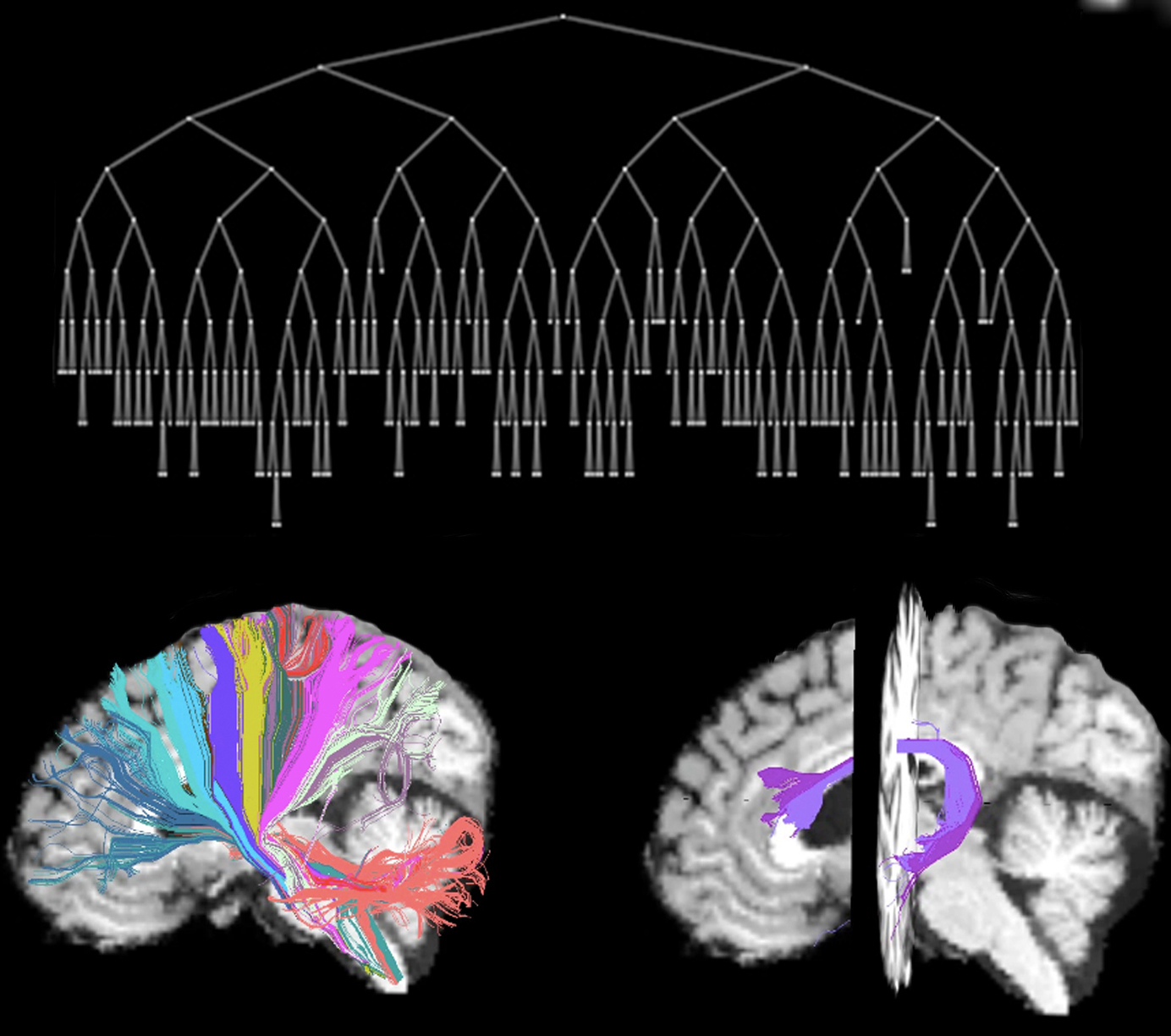
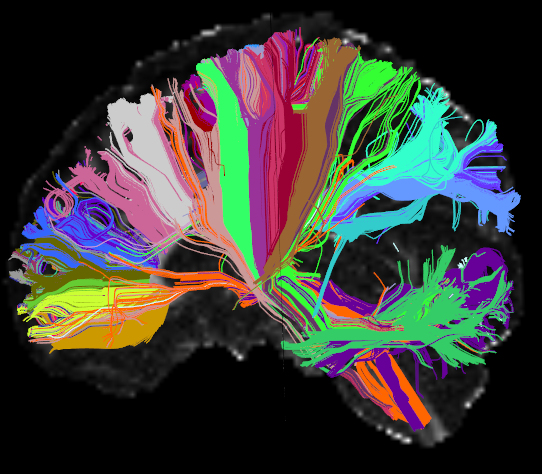
| = AnatomiCuts = | '''AnatomiCuts''' is an unsupervised hierarchical clustering that uses an anatomical similarity metric to cluster together white matter streamline with similar neighboring anatomical structures. |
| Line 3: | Line 3: |
| '''AnatomiCuts''' is an unsupervised hierarchical clustering that uses an anatomical similarity metric. | |
| Line 8: | Line 7: |
| == Inputs == - streamllines file (*.trk) - a segmentation image, preferable including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc.mgz or wm2009parc.mgz). |
= AnatomiCuts stream = |
| Line 12: | Line 9: |
| It is important that both files are in the same space. To validate this you can open the streamline file and the segmentation using both Freeview and Trackvis, since Freeview does not take into account RAS orientation, and Trackvis does not take into account the image offset. | == Enviroment variables == First you will need to set your SUBJECTS_DIR. This directoy should contain your recon outputs (mri folder). It is also necesary the dmri_preproc output (dmri folder). If this command was run on another directory, you can specify it with the DMRI_DIR variable. Additionally, you can specify a different output directory by setting ODMRI_DIR. By default, everything will point to your SUBJECTS_DIR. {{{ setenv SUBJECTS_DIR "dir with recon folders (mri)" }}} Optionally, you can have dmri_preproc and dmri.ac in different folders but setting the following variables: {{{ setenv DMRI_DIR "dir with mri_preproc outputs (dmri)" setenv ODMRI_DIR "output dir for dmri.ac (anatomicuts)" }}} == Deterministic tractography == For running deterministic tractography of GQI you would need to run the following command: {{{ dmri_ac tractography SUBJECT_ID }}} |
| Line 15: | Line 31: |
| Optionally, if you are in the center, to run it on the cluster you could run it like this: {{{ dmri_ac forAll tractography - "pbsubmit -c " }}} == Microstructural MAP (DTI/DKI) == You can extract DTI and/or DKI maps as well. However, for DKI you will need to have more than 1 bvalue in your data. {{{ dmri_ac getMaps SUBJECT_ID DTI dmri_ac getMaps SUBJECT_ID DKI }}} Optionally, if you are in the center, to run it on the cluster you could run it like this: {{{ dmri_ac forAll getMaps DTI "pbsubmit -c " dmri_ac forAll getMaps DKI "pbsubmit -c " }}} == Run == = AnatomiCuts executable = == Inputs == - streamlines file (*.trk) - segmentation image, ideally, including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc or wm2009parc). '''It is important to ensure that the white matter streamlines and the segmentation are in the same space.''' |
|
| Line 16: | Line 65: |
| Sometimes deterministic tractography can generate spurious streamlines outside the brain. To exclude them we can use [[streamlineFilter| streamlineFilter]]. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines. | Sometimes deterministic tractography can generate spurious streamlines outside the brain, or very short cut off streamlines. To exclude them we can use [[streamlineFilter| streamlineFilter]]. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines. |
| Line 18: | Line 67: |
| = Running AnatomiCuts = | == Running AnatomiCuts = |
| Line 28: | Line 77: |
| -c number of clusters -s segmentation file such as wm2009parc.nii.gz -f streamline file in trk format. |
|
| Line 36: | Line 79: |
| -n number of points to be used per streamline. The default is 10. | -n number of points to be sampled per streamline. The default is 10. |
| Line 38: | Line 81: |
| -e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice less than 300 is not recommended. | -e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice, less than 300 is not recommended. |
| Line 40: | Line 83: |
| -d directional neighbors to be used, the default is "a" all (26), diagonal "d" is 14 and straight "s" is 6. | -d directional neighbors to be used, the default is "a" all with 26 directions, diagonal "d" contains 14 directions and straight "s" only 6 directions. |
| Line 42: | Line 85: |
| -n number of points to be used per streamline. The default is 10. | -o output folder |
| Line 44: | Line 87: |
| -e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice less than 300 is not recommended (See Siless et al, 2018). | = Finding corresponding clusters across subjects = |
| Line 46: | Line 89: |
| -d directional neighbors to be used, the default is "a" all (26), diagonal "d" is 14 and straight "s" is 6. (See Siless et al 2018). | The Hungarian algorithm allows to find one-to-one correspondences and it's implementation for AnatomiCuts at multiple levels of the tree hierarchy can be found in [[AnatomiCuts_correspondences| AnatomiCuts_correspondences]]. |
| Line 48: | Line 91: |
| = Visualizing AnatomiCuts in Freeview = | |
| Line 49: | Line 93: |
| To load the clusters obtained with AnatomiCuts go to "File -> Load Tract Cluster" and select the AnatomiCuts output folder. | |
| Line 50: | Line 95: |
| {{attachment:anatomicuts.gif}} | |
| Line 55: | Line 99: |
V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.” |
AnatomiCuts is an unsupervised hierarchical clustering that uses an anatomical similarity metric to cluster together white matter streamline with similar neighboring anatomical structures.
Contents
AnatomiCuts stream
Enviroment variables
First you will need to set your SUBJECTS_DIR. This directoy should contain your recon outputs (mri folder). It is also necesary the dmri_preproc output (dmri folder). If this command was run on another directory, you can specify it with the DMRI_DIR variable. Additionally, you can specify a different output directory by setting ODMRI_DIR. By default, everything will point to your SUBJECTS_DIR.
setenv SUBJECTS_DIR "dir with recon folders (mri)"
Optionally, you can have dmri_preproc and dmri.ac in different folders but setting the following variables:
setenv DMRI_DIR "dir with mri_preproc outputs (dmri)" setenv ODMRI_DIR "output dir for dmri.ac (anatomicuts)"
Deterministic tractography
For running deterministic tractography of GQI you would need to run the following command:
dmri_ac tractography SUBJECT_ID
Optionally, if you are in the center, to run it on the cluster you could run it like this:
dmri_ac forAll tractography - "pbsubmit -c "
Microstructural MAP (DTI/DKI)
You can extract DTI and/or DKI maps as well. However, for DKI you will need to have more than 1 bvalue in your data.
dmri_ac getMaps SUBJECT_ID DTI dmri_ac getMaps SUBJECT_ID DKI
Optionally, if you are in the center, to run it on the cluster you could run it like this:
dmri_ac forAll getMaps DTI "pbsubmit -c " dmri_ac forAll getMaps DKI "pbsubmit -c "
Run
AnatomiCuts executable
Inputs
- streamlines file (*.trk)
- segmentation image, ideally, including cortical and subcortical parcellation with white matter segmentation based on neighboring regions (wmparc or wm2009parc).
It is important to ensure that the white matter streamlines and the segmentation are in the same space.
Filtering streamlines
Sometimes deterministic tractography can generate spurious streamlines outside the brain, or very short cut off streamlines. To exclude them we can use streamlineFilter. It is recommended to remove the short streamlines that can prematurely end during tractography. It is also possible to remove the ushape streamlines.
== Running AnatomiCuts =
AnatomiCuts -s segmentation.nii.gz -f streamlines.vtk -labels -o outputFolder <options>
Where
-s segmentation file such as wm2009parc.nii.gz
-f streamline file in trk format.
-c number of clusters. The default is 200.
-n number of points to be sampled per streamline. The default is 10.
-e number of streamlines to be used for the eigendecomposition of normalized cuts algorithm. The default is 500. Using less number of streamlines will fasten substantially the algorithm at the expense of accuracy. For a quick test 50 could be ok, but in practice, less than 300 is not recommended.
-d directional neighbors to be used, the default is "a" all with 26 directions, diagonal "d" contains 14 directions and straight "s" only 6 directions.
-o output folder
Finding corresponding clusters across subjects
The Hungarian algorithm allows to find one-to-one correspondences and it's implementation for AnatomiCuts at multiple levels of the tree hierarchy can be found in AnatomiCuts_correspondences.
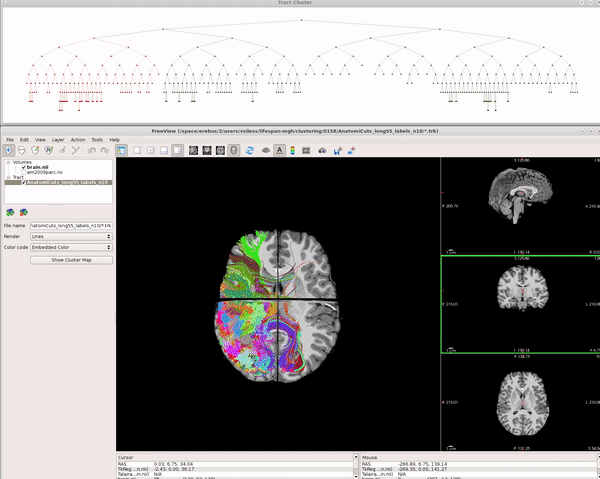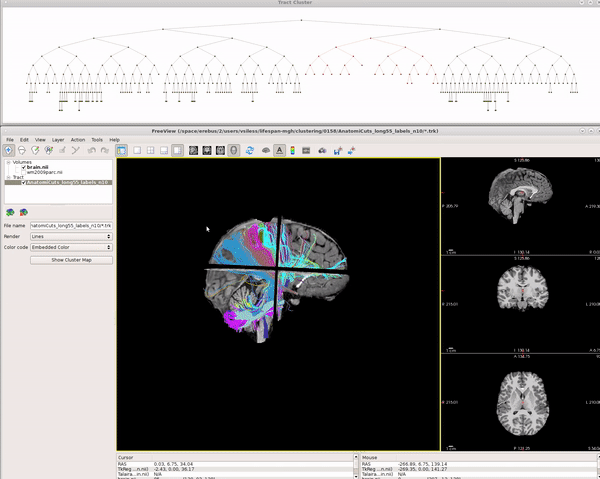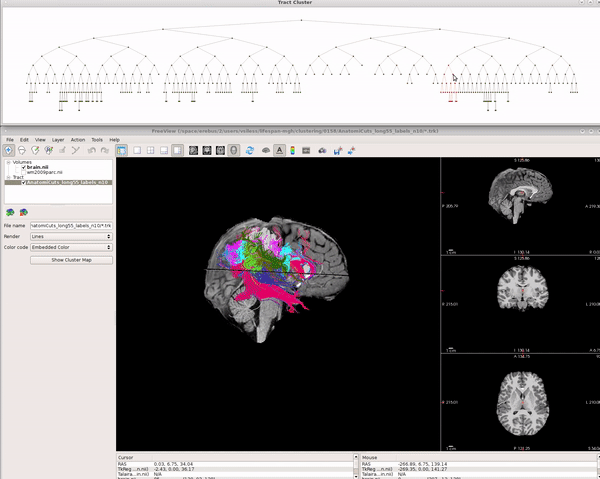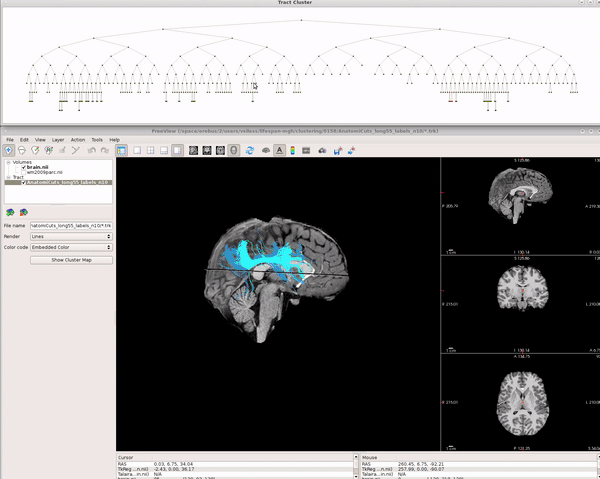
Visualizing AnatomiCuts in Freeview
To load the clusters obtained with AnatomiCuts go to "File -> Load Tract Cluster" and select the AnatomiCuts output folder.
References
AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity. Siless V., Chang K., Fischl B., Yendiki A.. NeuroImage 2018.
V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.”