| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| === work in progress... === | |
| Line 3: | Line 3: |
| Walkthrough: How to use FsFast and [[http://surfer.nmr.mgh.harvard.edu/fswiki/fcseed-sess|fcseed-sess]] for functional connectivity analysis including example commands. | This page describes how to perform seed-based functional connectivity (FC) analysis in FSFAST. This is an extension of the task-based analysis for which there is much more documentation. It may be worth your time to study some of the preprocessing and task-based analysis as found in [[http://surfer.nmr.mgh.harvard.edu/pub/docs/freesurfer.fsfast.ppt|FS-FAST powerpoint]] |
| Line 5: | Line 5: |
|
For general tips on using FsFast, download this [[http://surfer.nmr.mgh.harvard.edu/pub/docs/freesurfer.fsfast.ppt|FS-FAST powerpoint]] This walkthrough demonstrates how to run a functional connectivity analysis on resting state fMRI data. *STEP 1: Unpack Data into the FSFAST Hierarchy using [[https://surfer.nmr.mgh.harvard.edu/fswiki/unpacksdcmdir|unpacksdcmdir]] |
*STEP 1: Unpack Data into the FSFAST Hierarchy using dcmunpack (run with -help for more documentation): |
| Line 13: | Line 9: |
| unpacksdcmdir -src dicomdir/subject/ALLDICOMS -targ fcMRI_dir/subject -cfg subject_config.txt -fsfast -unpackerr | dcmunpack -src dicomdir/subject/ALLDICOMS -targ fcMRI_dir/subject -fsfast -run 3 bold nii.gz f.nii.gz -run 4 bold nii.gz f.nii.gz |
| Line 19: | Line 15: |
|
* subject_config.txt is a configuration text file you create (format below) * Use "-fsfast" to generate fsfast hierarchy subject_config.txt format: 28 bold nii f.nii 29 bold nii f.nii Col.1: scan acquisition number Col.2: output dir name will be created within "fcMRI_dir/subject" Col.3: output file format - this example is nifti format Col.4: output filename. In this example, 2 files will be created: . fcMRI_dir/subject/028/f.nii fcMRI_dir/subject/029/f.nii *QA Check after unpacking: * A - Check unpacked data (time points, # of slices ..etc) * B - Check FSFAST hierarchy in session folder *STEP 2: Reconstruction Anatomical data using [[https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all|recon-all]] Sample cmd: setenv SUBJECTS_DIR /path/to/recon_dir/ ; recon-all -s subject_dirname -all -i pathtoT1dicom_scan1.dcm -i pathtoT1dicom_scan2.dcm In this sample command... * set your SUBJECTS_DIR variable to your FreeSurfer subject recon directory * set the subject's directory name with "-s" ... the arguement you provide will become the directory name within $SUBJECTS_DIR * use "-i" to supply the dicoms to reconstruct. Use one "-i" per T1 acquisition. A. QA Check: * A - Check talairach transformation * B - Check skull strip, white matter & pial surface * C - Re-run "recon-all" if edits are made * D - Check hierarchy of reconstructed anatomical data B. Use FSFAST directory hierarchy: |
* -run 3 bold nii.gz f.nii.gz will unpack run 3 fmri to fcMRI_dir/subject/bold/003/f.nii.gz * To get a list of runs, run dcmunpack -src dicomdir/subject/ALLDICOMS * Use "-fsfast" to generate fsfast hierarchy shown in the image below |
| Line 61: | Line 21: |
| C. Link to FreeSurfer anatomical analysis: Create "subjectname" text file in the session directory. Write in it the subject's recon directory name (as labeld in $SUBJECTS_DIR). | *STEP 2: Link to FreeSurfer anatomical analysis: Create "subjectname" text file in the session directory. Write in it the subject's recon directory name (as labeld in $SUBJECTS_DIR). |
| Line 63: | Line 23: |
| D. Create a sessid file (text file with list of your sessions) in your Study DIR (optional) | C. Create a sessid file (text file with list of your sessions) in your Study DIR (optional) |
About
This page describes how to perform seed-based functional connectivity (FC) analysis in FSFAST. This is an extension of the task-based analysis for which there is much more documentation. It may be worth your time to study some of the preprocessing and task-based analysis as found in FS-FAST powerpoint
*STEP 1: Unpack Data into the FSFAST Hierarchy using dcmunpack (run with -help for more documentation):
Sample cmd:
dcmunpack -src dicomdir/subject/ALLDICOMS -targ fcMRI_dir/subject -fsfast -run 3 bold nii.gz f.nii.gz -run 4 bold nii.gz f.nii.gz
In this sample command...
- Have all fMRI dicoms linked into "ALLDICOMS" directory
- Arguement for "-targ" specifies output directory
- -run 3 bold nii.gz f.nii.gz will unpack run 3 fmri to fcMRI_dir/subject/bold/003/f.nii.gz
- To get a list of runs, run dcmunpack -src dicomdir/subject/ALLDICOMS
- Use "-fsfast" to generate fsfast hierarchy shown in the image below
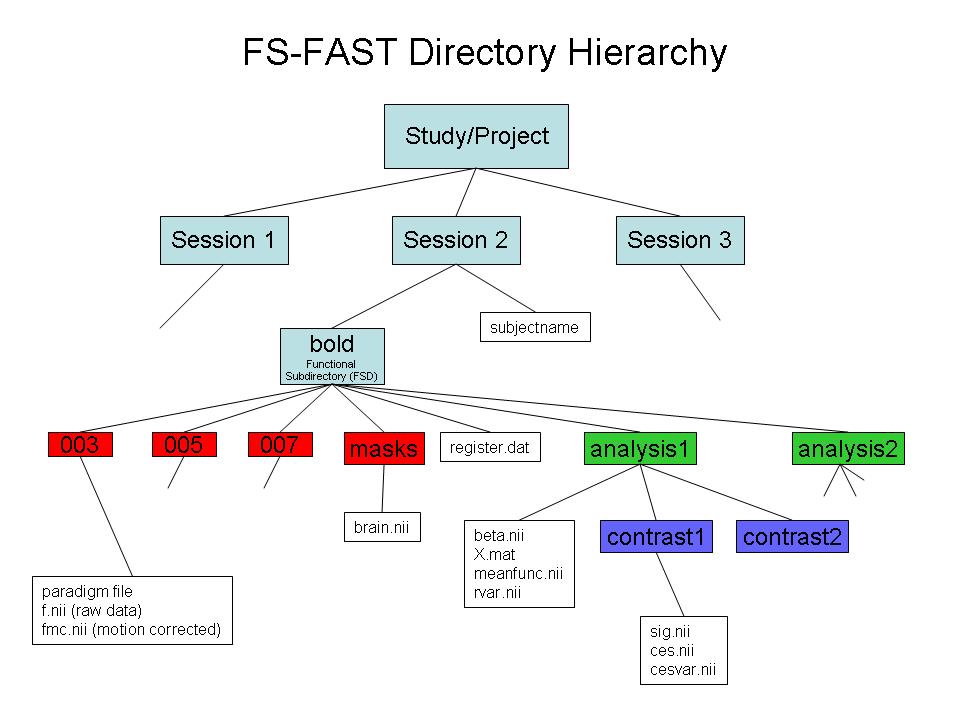
*STEP 2: Link to FreeSurfer anatomical analysis: Create "subjectname" text file in the session directory. Write in it the subject's recon directory name (as labeld in $SUBJECTS_DIR).
C. Create a sessid file (text file with list of your sessions) in your Study DIR (optional)
*STEP 3: Pre-process your bold data using preproc-sess preproc-sess
Sample cmd:
preproc-sess -s <subjid> -fwhm <#>
A. By default this will do motion correction, smoothing & brain masking
B. Quality Check (plot-twf-sess)
C.Examine additions to FSFAST hierarchy (in each run of bold dir):
f.nii
(Raw fMRI data)
fmc.nii
(Motion corrected-MC)
fmcsm5.nii
(MC & smoothed)
fmc.mcdat
(Text file with the MC parameters (AFNI))
brain.mgz
(Binary mask of the brain)
NOTE: you may need to convert the file "fmcpr.mgz" to fmcpr.nii using mri_convert
Found in each bold scan dir. Sample cmd:
mri_convert session/bold/002/fmcpr.mgz session/bold/002/fmcpr.nii
mri_convert session/bold/003/fmcpr.mgz session/bold/003/fmcpr.nii
Next, I'll outline two methods of deriving a seed region:
1) To use a full-size Freesurfer parcellation from aparc+aseg.mgz, continue with STEP 4 on this page.
2) To split the full Freesurfer parcellation into multiple seeds ("split parcellation"), follow the additional steps here - and resume with step 5 on this page...
*STEP 4: Use fcseed-config to record the parameters you wish to pass to your connectivity analysis.
Sample command: fcseed-config -segid 1010 -fcname mean.L_Posteriorcingulate.dat -fsd bold -mean -cfg mean.L_Posteriorcingulate.config
This example will use the FreeSurfer cortical segmentation for the left posterior cingulate (segID: 1010). For seed regions, we recommend generating the mean signal timecourse by using "-mean"
*STEP 5: Pass the config text file to fcseed-sess to generate time-course information for your chosen seed region (or for nuisance variable signal).
Sample cmd (mean seed region timecourse):
fcseed-sess -s <session> -cfg mean.L_Posteriorcingulate.config
Sample cmd (mean waveforms for nuisance regressors, but PCA also possible with -pca):
for white matter:
- fcseed-config -wm -fcname wm.dat -fsd bold -cfg wm.config
fcseed-sess -s <session> -cfg wm.config
for ventricles + CSF:
- fcseed-config -vcsf -fcname vcsf.dat -fsd bold -mean -cfg vcsf.config
fcseed-sess -s <session> -cfg vcsf.config
- NOTE: Once a config file is created it may be used for multiple sessions
*STEP 5: Use mkanalysis-sess to setup an analysis for your FC data
Sample cmd:
mkanalysis-sess -a <analysisname>
-surface fsaverage <hemi> -notask -taskreg mean.L_Posteriorcingulate.dat 1 -nuisreg vcsfreg.dat 1 -nuisreg wmreg.dat 1 -nuisreg global.waveform.dat 1 -fwhm 5 -fsd bold -TR <TR> -mcextreg -polyfit 5 -nskip 4
Note: if you do not want to regress out the global signal, then do not include it in the mkanalysis-sess command.
*STEP 6: Use selxavg3-sess to run the subject-level analysis outlined by the above mkanalysis-sess cmd.
selxavg3-sess -s <session> -a <analysisname>
*STEP 7: Choose the contrast file (generated in each session's contrast directory) that you wish to analyze on a group level:
- # ces.mgz - contrast effect size (contrast matrix * regression coef) # cesvar.mgz - variance of contrast effect size # sig.mgz - significance map (-log10(p)) # pcc.mgz - partial correlation coefficient map
*STEP 8: To continue with a group-level analysis, try one of the methods below:
- Method 1:
- create fsgd file containing all sessions of interest
Concatenate contrast files using mri_concat
Run group analysis using mri_glmfit
Concatenate with isxconcat-sess
Run group analysis using mri_glmfit
