|
Size: 4543
Comment:
|
Size: 5048
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| [[FsTutorial|top]] | [[FsTutorial/FslFeatFreeSurfer|previous]] | [[https://surfer.nmr.mgh.harvard.edu/fswiki/Tutorials|Back to list of all tutorials]]<<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial|Back to course page]]<<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/FslFeatFreeSurfer|FslFeat Tutorial Homepage]] ---- |
| Line 31: | Line 33: |
| Line 33: | Line 36: |
| The cortical and subcortical segmentations automatically generated by freesurfer can be mapped into the functional space, which can be useful for doing region of interest (ROI) analysis. This can be done with aseg2feat: | |
| Line 34: | Line 38: |
| The cortical and subcortical segmentations automatically generated by freesurfer can be mapped into the functional space, which can be useful for doing region of interest (ROI) analysis. This can be done with aseg2feat: | |
| Line 43: | Line 46: |
| The segmentation for a particular structure can be extracted to create a binary mask (i.e., a volume where the voxel value is 1 if it is in the structure and 0 otherwise). To make a binary mask of the left putamen, which has been assigned label 12 (see ${FREESURFER_HOME}/FreeSurferColorLUT.txt), use the following command: | |
| Line 44: | Line 48: |
| The segmentation for a particular structure can be extracted to create a binary mask (i.e., a volume where the voxel value is 1 if it is in the structure and 0 otherwise). To make a binary mask of the left putamen, which has been assigned label 12 (see ${FREESURFER_HOME}/FreeSurferColorLUT.txt), use the following command: | |
| Line 53: | Line 56: |
| Line 58: | Line 62: |
| -fthresh 0.5 -segmentation aparc+aseg.mgz | -fthresh 0.5 -segmentation aparc+aseg.mgz |
| Line 61: | Line 65: |
| You should see the image below:<<BR>> {{attachment:tkm-lh.putamen-cor-128-small.jpg}} |
You should see the image below:<<BR>> {{attachment:tkm-lh.putamen-cor-128-small.jpg}} |
| Line 66: | Line 69: |
| Line 73: | Line 77: |
| This will create fbert1.segstats.dat which will be a text file with a table of data. Each row will be a segmentation. The columns will contain various measures, including the number of functional voxels and the mean, stddev, min, max, and range of the cope over each ROI. [[FsTutorial/FslFeatSegStats| Here is an example]]. | This will create fbert1.segstats.dat which will be a text file with a table of data. Each row will be a segmentation. The columns will contain various measures, including the number of functional voxels and the mean, stddev, min, max, and range of the cope over each ROI. [[FsTutorial/FslFeatSegStats|Here is an example]]. ---- [[https://surfer.nmr.mgh.harvard.edu/fswiki/Tutorials|Back to list of all tutorials]]<<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial|Back to course page]]<<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/FslFeatFreeSurfer|FslFeat Tutorial Homepage]] |
Back to list of all tutorials
Back to course page
FslFeat Tutorial Homepage
Mapping automatic segmentations to the functional space
Preparations
If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/fbert-feat ln -s $FREESURFER_HOME/subjects/fsaverage ln -s $FREESURFER_HOME/subjects/bert
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window (this will paste the command). Press enter to run the command.
These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into this directory. You can now skip ahead to the tutorial (below the gray line).
If You're not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
tcsh source your_freesurfer_dir/SetUpFreeSurfer.csh setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/fbert-feat ln -s $FREESURFER_HOME/subjects/fsaverage ln -s $FREESURFER_HOME/subjects/bert
Notice the command to open tcsh. If you are already running the tcsh command shell, then the 'tcsh' command is not necessary. If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
The subjects 'fsaverage' and 'bert' may already exist in the subjects directory, so ignore any warnings that might be issued that say 'File exists'.
1.0 Mapping the segmentations
The cortical and subcortical segmentations automatically generated by freesurfer can be mapped into the functional space, which can be useful for doing region of interest (ROI) analysis. This can be done with aseg2feat:
aseg2feat --feat fbert1.feat --aseg aparc+aseg
This command will create fbert1.feat/reg/freesurfer/aparc+aseg.nii.gz. These are segmentations, meaning that each voxel has an integer value that corresponds to a particular structure. The mapping from structure number to name is given in ${FREESURFER_HOME}/FreeSurferColorLUT.txt.
2.0 Creating binary masks
The segmentation for a particular structure can be extracted to create a binary mask (i.e., a volume where the voxel value is 1 if it is in the structure and 0 otherwise). To make a binary mask of the left putamen, which has been assigned label 12 (see ${FREESURFER_HOME}/FreeSurferColorLUT.txt), use the following command:
fslmaths ./fbert1.feat/reg/freesurfer/aparc+aseg.nii.gz \
-thr 12 -uthr 12 \
./fbert1.feat/reg/freesurfer/lh.putamen.nii.gzTo view this binary mask on the anatomical:
tkmedit bert orig.mgz -aux brain.mgz \
-overlay ./fbert1.feat/reg/freesurfer/lh.putamen.nii.gz \
-overlay-reg ./fbert1.feat/reg/freesurfer/anat2exf.register.dat \
-fthresh 0.5 -segmentation aparc+aseg.mgz You should see the image below:
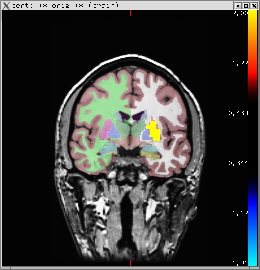
3.0 Creating ROI summaries
Once you have the segmentation mapped to the subject's native functional space, you can create summaries of the functional activation. Eg,:
mri_segstats --seg fbert1.feat/reg/freesurfer/aparc+aseg.nii.gz \ --nonempty --ctab-default \ --in fbert1.feat/stats/cope1.nii.gz --sum fbert1.segstats.dat
This will create fbert1.segstats.dat which will be a text file with a table of data. Each row will be a segmentation. The columns will contain various measures, including the number of functional voxels and the mean, stddev, min, max, and range of the cope over each ROI. Here is an example.
Back to list of all tutorials
Back to course page
FslFeat Tutorial Homepage
