Fixing a bad skull strip
To follow this exercise exactly, be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names.
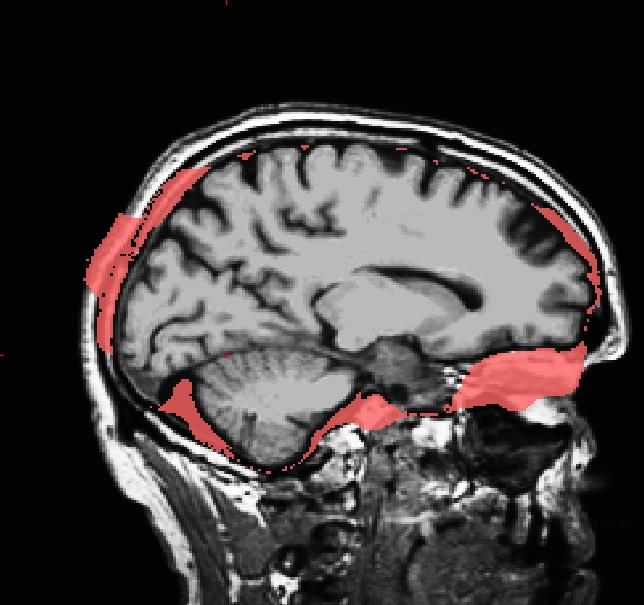
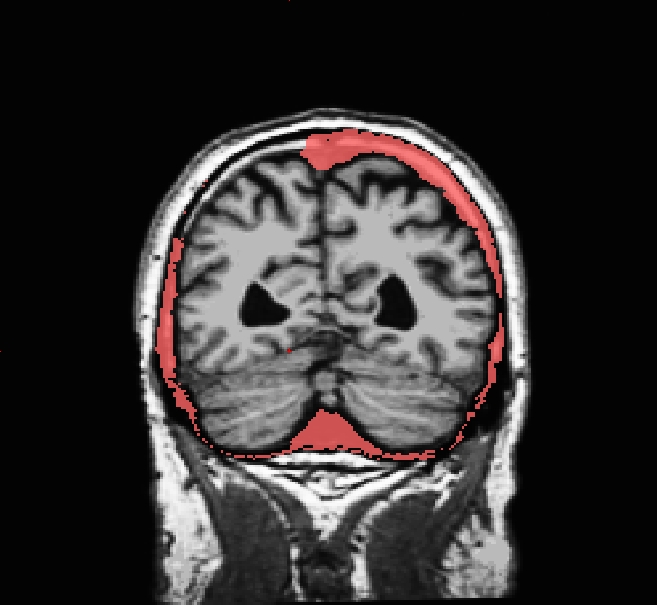
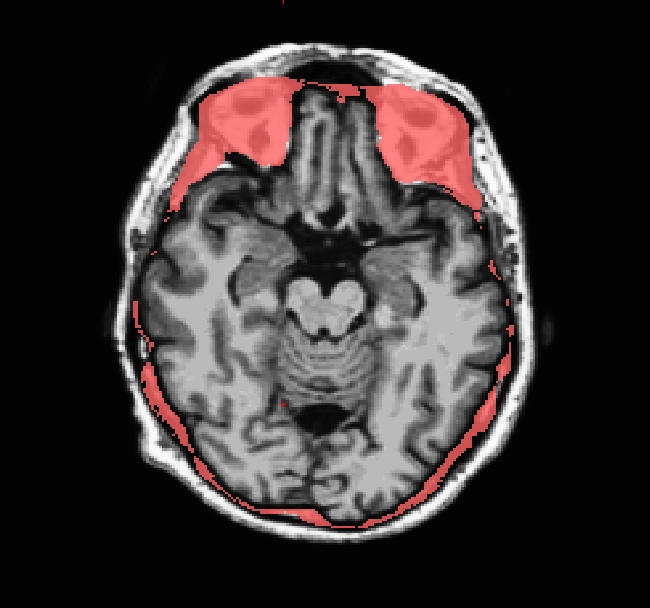
Occasionally, the skull stripping step removes more than just the skull, causing parts of the cortex or cerebellum to be removed. The skull stripping step can also fail to remove enough skull, leaving behind portions of the skull. In most cases both of these problems need to be corrected before continuing to the next step, either by manually editing the volumes or by adjusting input parameters to the skull stripping step and then running the skull strip again until a good result is obtained. However, if portions of the skull are left behind but they do not affect the pial or white surface (as the image below suggests), this is not considered to be an error. The red arrow indicates a portion of dura and skull that were not removed during the skull stripping step. Often the sagittal view reveals skull strip failures. Note that the inflated 3D surface is a less reliable gauge of skull strip failure unless large portions of the brain are missing, or lots of skull is retained.
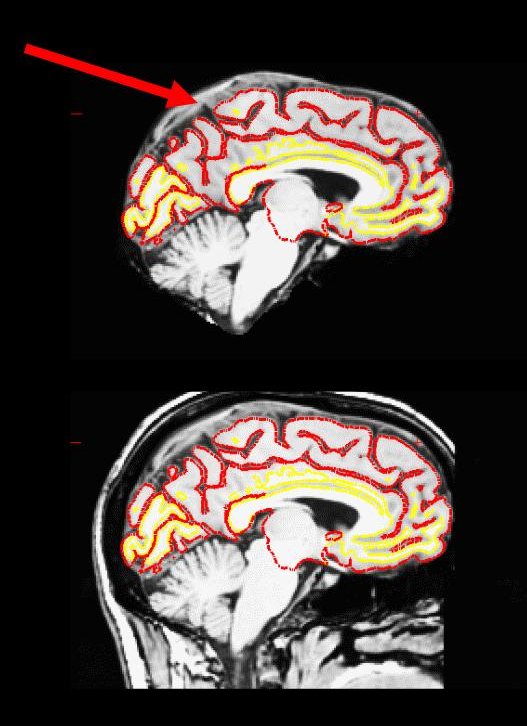
This page will walk you through the process of fixing a subject with a skull stripping problem. The subject below, skullstrip_before, has a poor skull strip in which posterior regions of the cortex have been stripped away along with the skull.
freeview -v skullstrip_before/mri/T1.mgz \ skullstrip_before/mri/brainmask.mgz:colormap=heat:visible=false \ skullstrip_before/mri/aparc+aseg.mgz:colormap=lut:opacity=0.2 \ -f skullstrip_before/surf/lh.white:edgecolor=yellow \ skullstrip_before/surf/lh.pial:edgecolor=red \ skullstrip_before/surf/rh.white:edgecolor=yellow \ skullstrip_before/surf/rh.pial:edgecolor=red
You can see that the segmentation and surfaces exclude some cortex in sagittal slice 101 (first image), and if you show the heat-mapped brainmask over the T1 volume (second image), you will see that these issues stem from a bad skull strip.
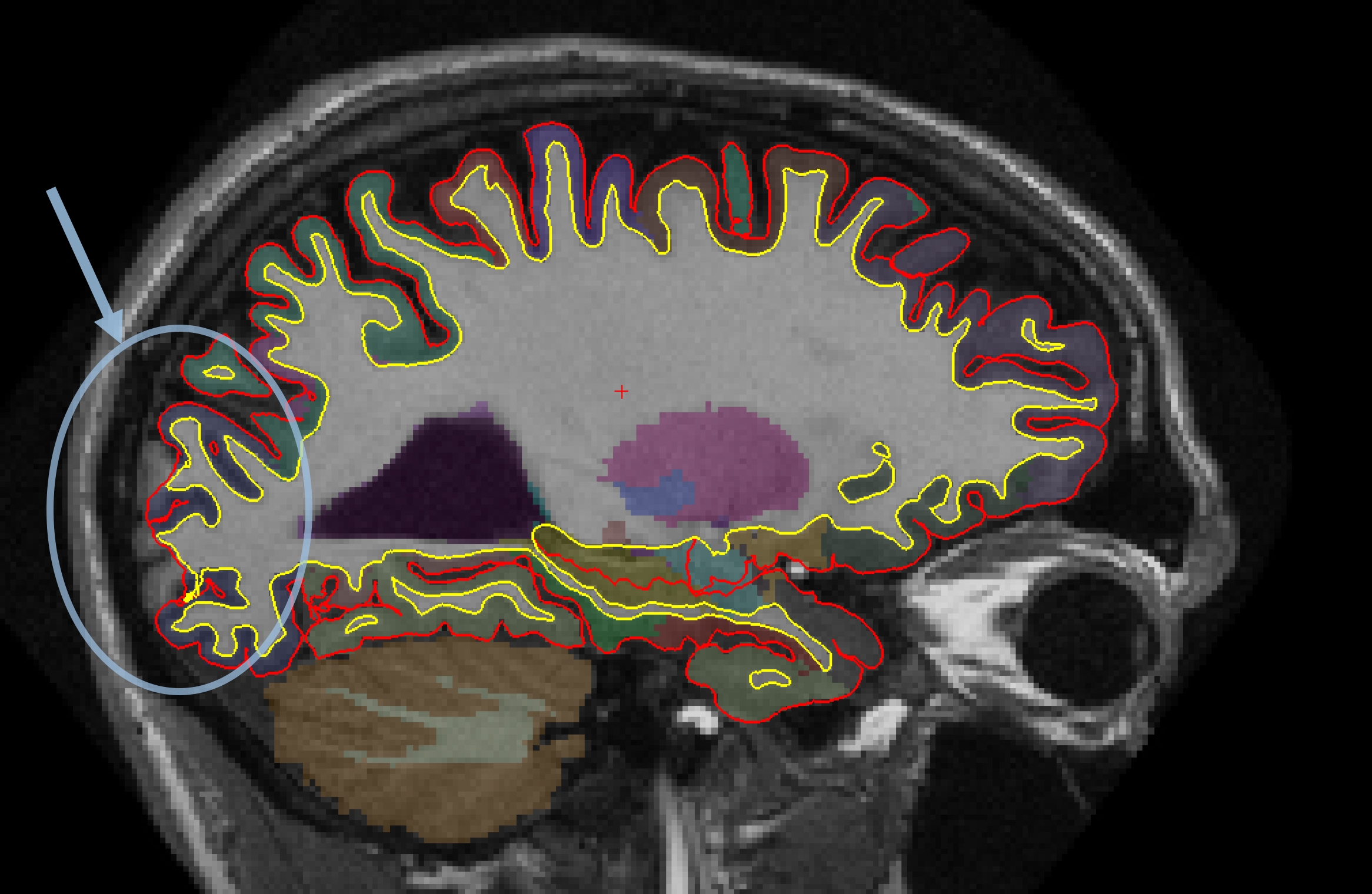
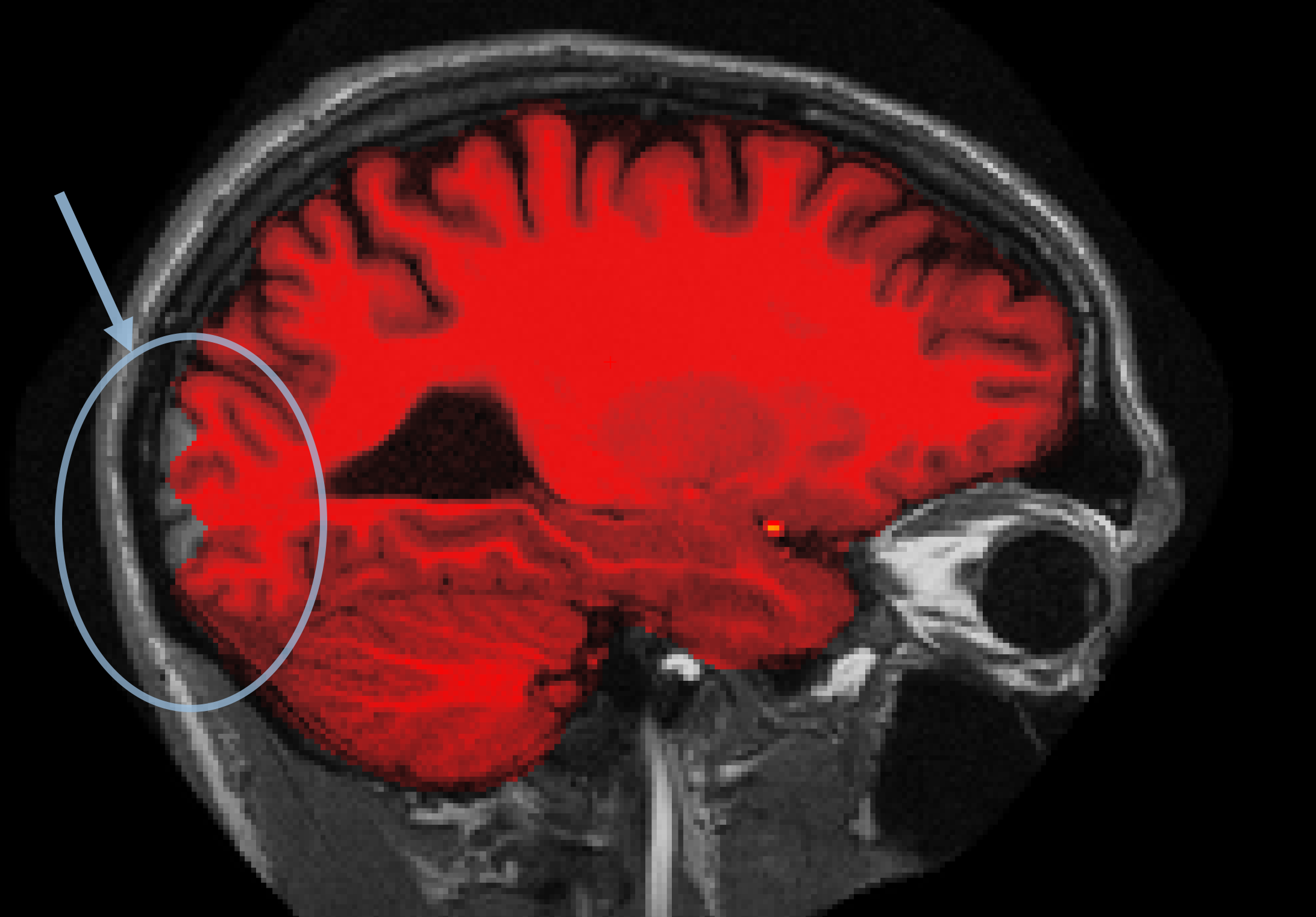
In general, there are two ways to fix a volume when there is something missing from the cortex or cerebellum - you can either clone in the missing pieces manually or adjust the parameters of mri_watershed to do it automatically. For this case, the easiest fix is to adjust the parameters of mri_watershed.
Adjusting watershed parameters
The watershed algorithm is used during the skull stripping step to find a boundary between the brain and skull. If you want the algorithm to be more conservative (i.e. if part of the brain has been removed), you will want to increase the watershed preflooding height percentage. If you want the algorithm to be more aggressive (i.e. part of the skull has been left behind), you will want to decrease the height. You can adjust the preflooding height by passing the following flags to recon-all:
recon-all -skullstrip -wsthresh <h> -clean-bm -subjid <subject name>
where <h> is replaced with the preflooding height you'd like to use and <subject name> is replaced with your subject. The clean-bm flag is used to instruct recon-all to write over the old brainmask.mgz volume with your new edits. If you do not use this flag, your changes will not take effect.
Part of the brain is missing
Now we will take another look at the above problem case, where part of the cortex was removed. If you suspect that certain anatomy could be an outlier, and may be responsible for making the skullstripping step fail or produce poor results, then you can try using the -no-wsgcaatlas flag (wsgcaatlas = with skull gaussian classifier array atlas). For this case, try fixing the brainmask by running the watershed command with a few different preflooding heights between 1 and 50:
recon-all -skullstrip -wsthresh <h> -clean-bm -no-wsgcaatlas -subjid skullstrip_before
Take a look at your output volume (brainmask.mgz has been changed) along with the original T1 volume (T1.mgz).
freeview -v skullstrip_after/mri/T1.mgz \ skullstrip_after/mri/brainmask.mgz:colormap=heat
The accurately skull-stripped brainmask should look something like this (sagittal slice 101):
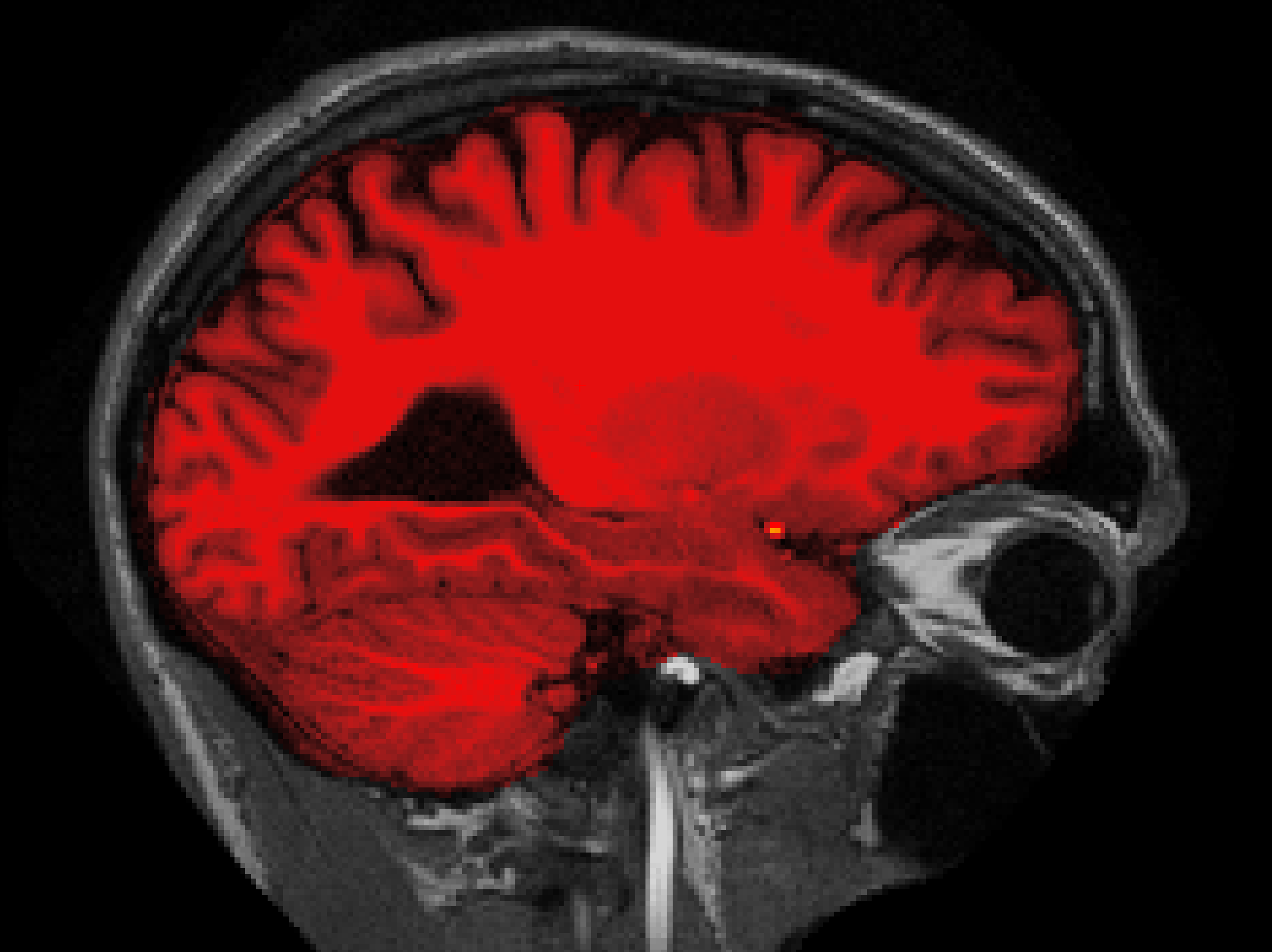
What watershed threshold value did you use to produce an optimal skull strip? Was is between 20 and 30? If so, that's because the default preflooding height is 25, which produces the cleanest skull strip for most cases. There aren't any hard and fast rules about how to select your height value, but as a general rule of thumb, if part of the brain is missing, you should start with a watershed threshold around 35, and if too much skull is remaining, you should start with a threshold of around 15.
Using gcut
When the skull stripping has left a small part of dura left, it is quicker to try and rerun the skull strip step using the -gcut flag than to do manual editing. This flag should automatically remove any extra dura that could influence the surfaces.
recon-all -skullstrip -clean-bm -gcut -subjid <subject name>
This flag runs the mri_gcut command. Please see this page or run mri_gcut --help for more information.
INFO: Care must be taken to thoroughly inspect your data when using -gcut. In particular, inspect the edges of gm and cerebellum for over-aggressive cutting. Open the brainmask.gcuts.mgz volume in freeview to view the voxels which gcut has removed. We recommend viewing the brainmask.gcuts.mgz overlayed on the T1.mgz to clearly see which voxels have been removed by gcut.
Regenerating surfaces after the bad skull strip is fixed
After the bad skull strip has been improved, you can run the following command to regenerate the final white and pial surfaces based on the new brainmask.mgz:
recon-all -autorecon-pial -subjid <subject name>
This command will not regenerate the aseg.mgz or wm.mgz files because those files are created earlier in the recon-all processing stream, prior to the point where surfaces are generated using mris_make_surfaces. For full steps please see: ReconAllTableStableV6.0. If you want to regenerate these files you must re-run from the beginning. To do so, run:
recon-all -all -subjid <subject name>
Do not run these commands if you are conducting the tutorial!
