| Deletions are marked like this. | Additions are marked like this. |
| Line 92: | Line 92: |
| * Volumes are saved under <subjects_directory>/<subject_name>/mri/hypothalamic_subunits_volumes.csv. |
* Volumes are saved under <subjects_directory>/<subject_name>/mri/hypothalamic_subunits_volumes.v1.csv. * Volumetric stats are saved under <subjects_directory>/<subject_name>/stats/hypothalamic_subunits_volumes.v1.stats. |
Segmentation of hypothalamic subunits
This functionality is available in the development version of FreeSurfer as of September 18th 2020.
Author: Benjamin Billot
E-mail: benjamin.billot.18 [at] ucl.ac.uk
Rather than directly contacting the author, please post your questions on this module to the FreeSurfer mailing list at freesurfer [at] nmr.mgh.harvard.edu
If you use these tools in your analysis, please cite:
Automated segmentation of the hypothalamus and associated subunits in brain MRI. Billot, B., Bocchetta, M., Todd, E., Dalca, A.V., Rohrer, J.D., Iglesias, J.E. Neuroimage (in press).
See also: ThalamicNuclei, HippocampalSubfieldsAndNucleiOfAmygdala, BrainstemSubstructures
Contents
- General Description
- Installation
- Usage
- Frequently asked questions
1. General Description
This tool enables automated segmentation of the hypothalamus and its associated subunits in 3D T1-weighted scans of approximately 1mm isotropic resolution. It uses a convolutional neural network, which enables computing segmentations in a very short processing time (around 10 seconds with a CPU, less than a second with a GPU). More specifically, this code produces segmentation maps for five subregions, which comprise the following hypothalamic nuclei:
Anterior-inferior: suprachiasmatic nucleus; supraoptic nucleus (SON)
Anterior-superior: preoptic area; paraventricular nucleus (PVN)
Posterior: mamillary body (including medial and lateral mamillary nuclei); lateral hypothalamus; tuberomamillary nucleus (TMN)
Inferior tubular: infundibular (or arcuate) nucleus; ventromedial nucleus; SON; lateral tubular nucleus; TMN
Superior tubular: dorsomedial nucleus; PVN; lateral hypothalamus
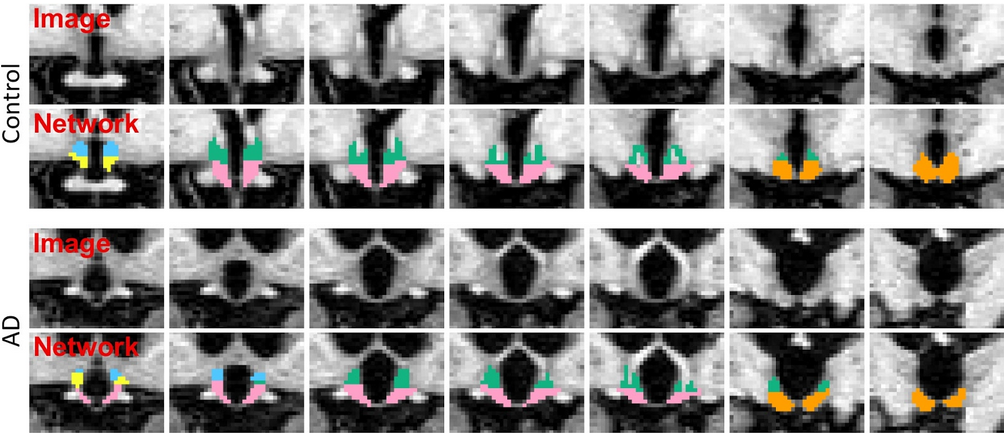
The segmentation model was trained by applying aggressive data augmentation, which makes it robust against variability in acquisition parameters (sequence, platform, bias field, head positioning), and in anatomy (e.g. atrophy patterns linked to ageing or different pathologies). The following figure illustrates two segmentation examples in coronal slices obtained by this model for a control subject, and a subject diagnosed with Alzheimer's disease:
2. Installation
The first time you run this module, it will prompt you to install Tensorflow. Simply follow the instructions in the screen to install the CPU or GPU version.
If you have a compatible GPU, you can install the GPU version for faster processing, but this requires installing libraries (GPU driver, Cuda, CuDNN). These libraries are generally required for a GPU, and are not specific for this tool. In fact you may have already installed them. In this case you can directly use this tool without taking any further actions, as the code will automatically run on your GPU.
3. Usage
This software can be used in two modes:
on subjects processed with FreeSurfer
- on any T1 scan of approximately 1mm resolution.
The first mode follows the FreeSurfer subject directory structure and works on the bias field corrected images (nu.mgz). We recommend using this mode if you are running the main FreeSurfer recon-all scripts on your data. The second mode is more flexible and has the advantage of not having to run subjects through recon-all, if you are only interested in the hypothalamus.
A summary of the following information can be found simply by typing:
mri_segment_hypothalamic_subunits -h
a. FreeSurfer subjects
This tool can be run on subjects that have been analyzed beforehand with the main FreeSurfer stream (i.e. “recon-all”). Therefore, we will assume that a command similar to the following one has already been run on each subject for which we want to segment the hypothalamic subunits:
recon-all -all --s <subject_name>
To analyze one or several subjects, simply use:
mri_segment_hypothalamic_subunits --s <subject_name> --sd <subjects_dir> --write_posteriors --threads <threads>
where:
--s <subject_name>: is the name of a subject in $SUBJECTS_DIR. You can also specify several subjects to run them all at the same time. If no name is given after --s, the code will be run on ALL subjects in $SUBJECTS_DIR.
--sd <subjects_dir>: (optional) is the path to a folder with which to override the current value of $SUBJECTS_DIR.
--write_posteriors: (optional) will save the posteriors (soft segmentations) obtained for each subject, in addition to the hard label map.
--threads <threads>: (optional) number of threads to be used by Tensorflow (default uses one core). Increase the number of threads to decrease the runtime when using the CPU version.
When running on a subject analysed with FreeSurfer, this software automatically saves the segmentation and volumes of the hypothalamic subunits in the FreeSurfer directory structure.
Segmentations are saved under <subjects_directory>/<subject_name>/mri/hypothalamic_subunits_seg.v1.mgz.
Volumes are saved under <subjects_directory>/<subject_name>/mri/hypothalamic_subunits_volumes.v1.csv.
Volumetric stats are saved under <subjects_directory>/<subject_name>/stats/hypothalamic_subunits_volumes.v1.stats.
Posteriors (when enabled) are saved under <subjects_directory>/<subject_name>/mri/hypothalamic_subunits_posteriors.v1.mgz.
Important: if several subjects are given, the outputs (segmentation, volumes, and possibly posteriors if requested) will be saved in each subject's directory. Additionally, a CSV file regrouping the volumes of all structures for all subjects will be saved under:
- $SUBJECTS_DIR/hypothalamic_subunits_volumes_all.v1.csv.
Important: If you wish to run this tool on several subjects, we highly recommend that you use a single call to mri_segment_hypothalamic_subunits in order to segment all the subjects in the subjects directory, rather than calling it once per subjects. This will run ***much*** faster as it avoids having to set up the software for each scan.
WARNING: we do not recommend to save the posteriors unless needed, as the process of writing these big files to disk considerably slows down the execution of the program.
Examples:
Segment all subjects in $SUBJECTS_DIR:
mri_segment_hypothalamic_subunits --s
Segment a single subject bert:
mri_segment_hypothalamic_subunits --s bert
Segment several subjects (bert and ernie):
mri_segment_hypothalamic_subunits --s bert ernie
Segment a single subject bert located in a subject directory other than $SUBJECTS_DIR:
mri_segment_hypothalamic_subunits --s bert --sd /home/user1/data/new_analysis/
Segment all subjects located in a subject directory other than $SUBJECTS_DIR:
mri_segment_hypothalamic_subunits --s --sd /home/user1/data/new_analysis/
b. Any T1 scan
You can also run this on a T1 scan that has not been processed with FreeSurfer. This tool works on scans that have been not subjects to any preprocessing (e.g., no bias field correction or skull stripping is needed). The command line is:
mri_segment_hypothalamic_subunits --i <input> --o <output> --out_posteriors <posteriors> --out_volumes <volume_file> --threads <threads>
where:
<input>: is the path to a single scan to segment. This can also be a folder, in which case all the scans in this folder will be segmented.
<output>: is the path where the output segmentation will be saved. This must be a file if <input> is a file, or a folder if <input> is a folder.
<posteriors>: (optional) is the path where the posteriors (given as soft probability maps) will be saved. This must also be a folder if <input> designates a folder.
<volume_file>: (optional) is the path to an output CSV file where the volumes of the whole hypothalamus and of all subunits will be saved. In the case where <input> designates a folder, the volumes of all subjects will be saved in the same file.
<threads>: (optional) number of threads to be used by Tensorflow (default uses one core). Increase the number of threads to decrease the runtime when using the CPU version.
In the case where volumes are saved, the corresponding file is organized as follows: each row corresponds to a different subject, and columns correspond to the different structures.
Important: If you wish to run this tool on several scans in the T1 mode, we highly recommend that you put them in a single folder and run mri_segment_hypothalamic_subunits on this folder, rather than calling it individually on each scan. This will enable you to save time as it avoids having to set up the software for each scan.
Examples:
Segment a single scan:
mri_segment_hypothalamic_subunits --i /home/user1/data/scan.mgz --o home/user1/data/seg.mgz
Segment a single scan and save the volumes:
mri_segment_hypothalamic_subunits --i /home/user1/data/scan.mgz --o home/user1/data/seg.mgz --out_volumes /home/user1/data/hypothalamic_volumes.csv
Segment all scans contained in a folder:
mri_segment_hypothalamic_subunits --i /home/user1/data/scans/ --o /home/user1/data/segmentations/
Segment all scans contained in a folder, and save the volumes:
mri_segment_hypothalamic_subunits --i /home/user1/data/scans/ --o /home/user1/data/segmentations/ --out_volumes /home/user1/data/hypothalamic_volumes.csv
4. Frequently asked questions (FAQ)
Does running this tool require preprocessing of the input scans?
No! Because we applied aggressive augmentation during training (see paper), this tool is able to segment both processed and unprocessed data. So there is no need to apply bias field correction, skull stripping, or intensity normalization.
The sum of the number of voxels of a given structure multiplied by the volume of a voxel is not equal to the volume reported in the output volume file.
This is because the volumes are computed upon a soft segmentation, rather than the discrete segmentation. The same happens with the main recon-all stream: if you compute volumes by counting voxels in aseg.mgz, you don't get the values reported in aseg.stats.
What formats are supported in the second mode (any T1 scan)?
This tool can be run on Nifti (.nii/.nii.gz) and FreeSurfer (.mgz) scans.
Which scan is actually used when inputting a FreeSurfer subject?
In this case, the module segments nu.mgz, which is corrected for bias field.
How can I increase the speed of the CPU version without using a GPU?
If you have a multi-core machine, you can increase the number of threads with the --threads flag (up to the number of cores).
