Contents
Introduction
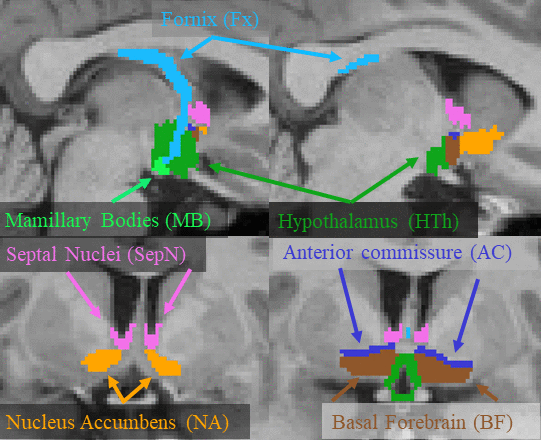
This page describes a deep learning tool, mri_sclimbic_seg, to automatically segment several subcorical limbic structures from T1-weighted images ("SC" = subcortical). The structures are shown in the figure below (note: anterior commissure is only present for reference reasons). Please cite: A Deep Learning Toolbox for Automatic Segmentation of Subcortical Limbic Structures from MRI Images Greve, DN, Billot, B, Cordero, D, Hoopes, M. Hoffmann, A, Dalca, A, Fischl, B, Iglesias, JE, Augustinack, JC. 2020. Submitted to Neuroimage.
Installation
Does not need the rest of FS.
Basic Usage
In the basic usage, one creates a folder with the T1 volumes in (NIFTI or mgz format) one wants to segment, then runs
mri_sclimbic_mri –i inputfolder –i outputfolder --write_volumes --write_qa_stats
The tool will find all the input images, segment them, and write out the segmentation volumes into the output folder. The inputs should be T1-weighted volumes. If the resolution is not 1mm3, then you must add the --conform flag to mri_sclimbic_seg. If the images are named subject1.mgz subject2.mgz, etc, the segmentations will be named subjec1.sclimbic.mgz, subject2.sclimbic.mgz, etc. The segmentation can be viewed with freeview or tkmeditfv:
freeview inputfolder/subject1.mgz outputvolder/subject1.sclimbic.mgz
and should look like the figure above.
The above mri_sc_limbic command will also create a CSV file where each row is a case, each column is a label, and each entry is the volume of that structure in mm3.
Quality Control
The z-score should only be used for quality control. It should not be interpreted as a deviation from a normal population because the subjects used for manual labeling were not chosen with this in mind.
Including Estimated Intracranial Volume (eTIV)
Group GLM Analysis
Running in FreeSurfer Mode
On a single threaded CPU, the program takes about 40sec to run on a single case; with 3 threads, the time drops to about 15sec; using a GPU does not reduce this significantly as much of the time is spent loading and writing. The tool uses about 20GB of memory. If the input volume is not 1mm3, then there is an option to reslice to this resolution; the reslicing is only internal – the output segmentation is resliced back to the original resolution. Note that this changing of resolution may affect the quality. If one is planning to perform a volumetric study, then one will need to normalize by ICV. If one does not have an estimate of the ICV, then the tool can compute it using the FreeSurfer method (Buckner et al., 2004); the ICV will be included as a column in the CSV file. Computing the ICV will increase the processing time to about 5min for each case. The CSV file can be imported into a statistical program like SPSS or R for further processing or it can be processed using FreeSurfer’s mri_glmfit, which includes automatic application of ICV correction if ICV is in the CSV. The user should visually spot check the segmentation output. To assist in quality control, the tool can output two more CSV files. One contains a z-score[1] for the volume each structure based on the means and standard deviations of the manual labels. In the other, the “confidence” (mean posterior probability within the label) is reported. If the z-score is very high or the confidence is very low, then the case should definitely be visually examined. The tool does not require knowledge of FreeSurfer; as long as FreeSurfer is installed properly, then the user need only understand and execute mri_sclimbic_seg.
[1] The z-score should only be used for quality control. It should not be interpreted as a deviation from a normal population because the subjects used for manual labeling were not chosen with this in mind.
