Research Overview
Mixture modelling is an alternative method of carrying out inference on (i.e., thresholding) statistical maps such as t, f or z stat images output by FEAT. Instead of just making assumptions about the null part of the statistical distribution and making all inference with respect to that, mixture modelling explicitly models the null part and the activation part. We can thus carry out "alternate hypothesis testing" and ask a richer set of questions of the data. Here we also implement a spatial enhancement to the mixture modelling; we use a spatial Markov random field to regularise (smooth) the labelling of voxels into null, activated or "de"-activated.
For more detail on the spatial mixture modelling see Mixture Models with Adaptive Spatial Regularisation for Segmentation with an Application to FMRI Data; Woolrich, M., Behrens, T., Beckmann, C., and Smith, S.; IEEE Trans. Medical Imaging, 24(1):1-11, 2005 and a related technical report TR04MW1.
mm
Warning: this implementation of spatial mixture modelling is a beta release and has not been extensively tested! To run spatial mixture modelling on a statistic image (such as grot.feat/stats/zstat1) use the mm command-line program. The minimum syntax is:
mm --mask=<mask_image> --sdf=<stats_image>
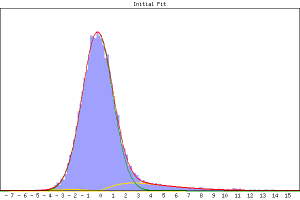
and the other main option that you might want to use is --zfstatmode which tells mm not to fit a deactivation class. Just type mm to get the full help output. The output from mm is summarised in a web-page named <outputdir>/MM.html and the probability of lying in each of the two or three segmentation classes (null, activation and de-activation) are found in <outputdir>/w1_mean, w2_mean and w3_mean. Note that the histogram plot in the web-page report will look less "nice" a fit than a non-spatial mixture model. This is because the final fitting is a balance between optimal histogram fitting of the class mixtures and the spatial processes which attempt to smooth voxel classification.
