The purpose of this tutorial is to give you experience with the integration of Diffusion Tensor Imaging (DTI) with FreeSurfer. The data were collected at MGH as part of the MIND project.
If you are looking for the Tracula tutorial, you can find it here.
DTI Basics
It is assumed that you already have some basic knowledge of DTI and its analysis. This paragraph is supplied as a summary for completeness. DTI attempts to measure the diffusion of water molecules in the tissue. Within the brain, white matter tracts tend to act like little straws that constrain the direction of diffusion. DTI analysis measures the orientation of this diffusion and the "strength" of this orientation. The strength of preferred orientation is often measured by fractional anisotropy (FA) and diffusivity is often measured by the Apparent Diffusion Coefficient (ADC).
Prior to DTI analysis, it is customary to motion correct, and sometimes eddy current correct, the diffusion weighted images. This is done by selecting one of the images as a template (usually a low-b volume). This forces all the DTI-related maps to be in-line with the template. Consequently, this template should be used as the registration target.
Tutorial Data Set
This data was acquired at MGH as part of a MIND Consortium study. Seventy images were acquired, 10 low-b and 60 diffusion weighted. The diffusion data are located in the '$TUTORIAL_DATA/diffusion_tutorial' directory and the corresponding FreeSurfer anatomical analysis (recon) is located in the '$TUTORIAL_DATA/diffusion_recons' directory.
Preparations
If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
export SUBJECTS_DIR=$TUTORIAL_DATA/diffusion_recons export TUTORIAL_DIR=$TUTORIAL_DATA/diffusion_tutorial subj=Diff001
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window (this will paste the command). Press enter to run the command.
These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into this directory. You can now skip ahead to the tutorial (below the gray line).
If You're not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
source your_freesurfer_dir/SetUpFreeSurfer.csh export TUTORIAL_DATA=your_tutorial_data export SUBJECTS_DIR=$TUTORIAL_DATA/diffusion_recons export TUTORIAL_DIR=$TUTORIAL_DATA/diffusion_tutorial subj=Diff001
If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
This tutorial uses Freeview to view the output on several processing steps. Click here to familiarize yourself with Freeview if you are not already.
DTI Data Analysis Processing Stream
Pre-processing
The following steps are necessary to prepare the diffusion data for various types of analyses. On each step there is a description of the command and how it relates to the overall processing stream, the actual command you would use, and how to visualize the output. Similar pre-processing steps can be accomplished by using Tracula, although output will be generated using different file names and file types. You may find that you prefer one method over the other depending on the types of analyses and degree of involvement you want to have.
DT_Recon
To get started, one would have to reconstruct the diffusion tensor data from images collected. Additionally, it is recommended to correct for eddy currents and any motion artifacts, which could distort the DTI data. The 'dt_recon' command can be used to accomplish this pre-processing. It generates multiple NIFTI volumes that will be used in later steps. By default, the 'dt_recon' command assumes that the FreeSurfer anatomical analysis (recon) has already been completed.
The following command is the first step in processing your diffusion data with dt_recon. If you are taking part of an organized course, DO NOT RUN THIS COMMAND. It would take a long time so it has already been run for you.
dt_recon --i $TUTORIAL_DIR/$subj/orig/*-1.dcm --s $subj --o $TUTORIAL_DIR/$subj/dtrecon
Notes:
- The subject we use as an example for this tutorial was scanned in a Siemens scanner. Thus, our input is in DICOM format. The bvalues and gradient directions are stored in the header and are written to bvals.dat and bvects.dat, respectively, during the following analysis. If your diffusion data was acquired using a non-Siemens scanner, you will have to specify the scan-specific bvalues and bvector with the '--b' flag when running the command above.
- The location of the DICOM files for our subject is indicated by the '--i' flag.
- In order to automatically establish a correspondence between the structural and diffusion scans of the same subject, we pass the subject ID information using the '--s' flag. That will allow the program to find the relevant files in '$SUBJECTS_DIR/$subj' of the structural recon. (If you do not have access to the structural recons at the time of the diffusion analysis, you can omit '--s' and use the '--no-reg' flag.)
- The desired location of the output is indicated with the '--o' flag. In this example, we are going to name the output folder 'dtrecon', which will be automatically created by the program.
To see a script which could be used to run dt_recon on multiple subjects take a look at the 'DiffPreproc.csh' script on this page.
Viewing the Data
After running the 'dt_recon' command, you may want to look at some of the outputs in Freeview. Run the following command to view the fractional anisotropy volume (fa.nii) and the apparent diffusion coefficient volume (adc.nii). The '$TUTORIAL_DIR' and '$subj' parameters need to be defined in advance. If you haven't done this already, please see the Tutorial Data Set section.
freeview -v $TUTORIAL_DIR/$subj/dtrecon/fa.nii \
$TUTORIAL_DIR/$subj/dtrecon/adc.niiScroll the middle mouse button to zoom in and out. Click and hold the middle mouse button to move around the image. Use the 'Page Up' and 'Page Down' keys to move back and forth through the volume slices. Use the orientation buttons to switch the plane (  ,
,  ,
,  ,
,  ). Toggle between the two volumes by using 'Alt + C' in the Volume Tab.
). Toggle between the two volumes by using 'Alt + C' in the Volume Tab.
Coronal view of the fractional anisotropy volume (fa.nii):
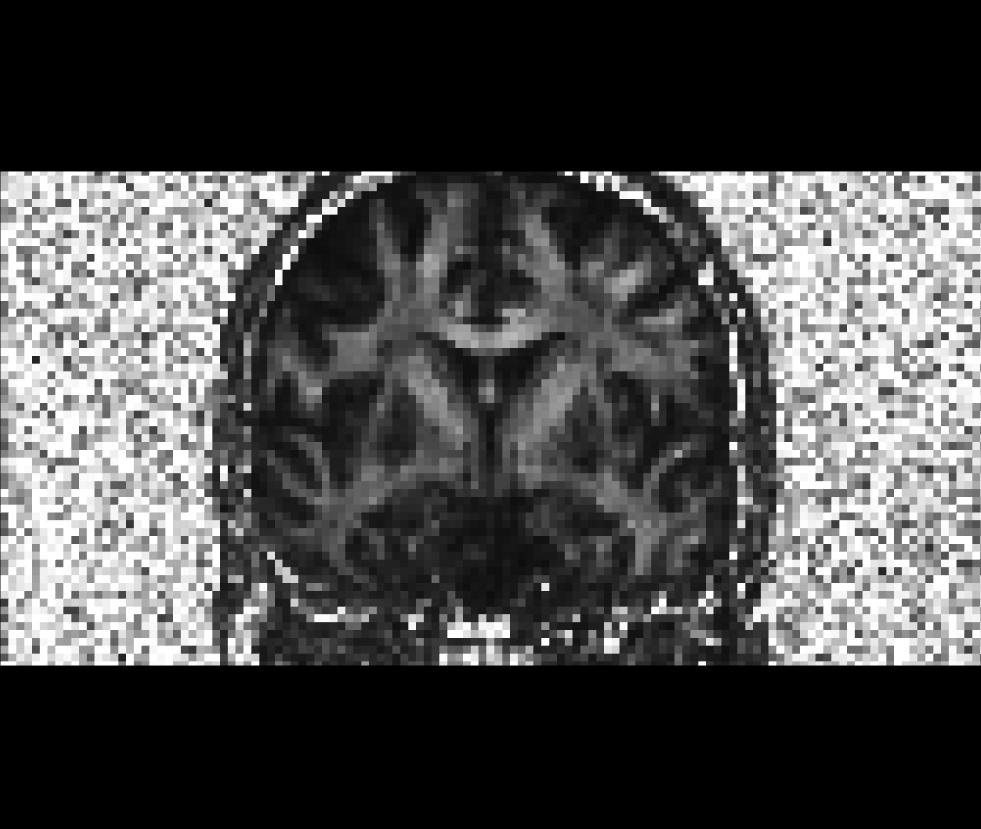
Coronal view of the apparent diffusion coefficient volume (adc.nii):
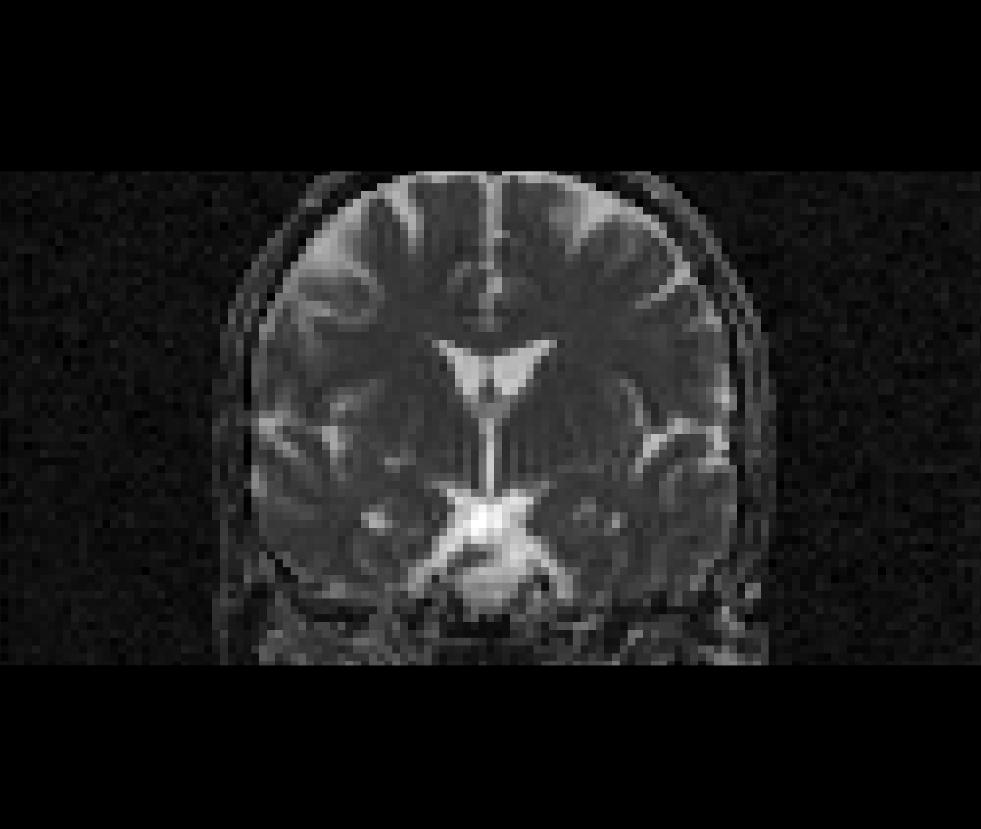
All other volumetric data sets generated by dt_recon (*.nii) may be viewed with Freeview. To see an explanation of the various outputs, see the dt_recon page. Close Freeview when you are done before proceeding to the next step.
Resampling Structural Data in the Diffusion Space
In order to analyze specific sub-cortical or white matter regions in the diffusion data, we can resample the aseg (subcortical segmentation created by recon-all) from the structural data into the diffusion space. The registration between the diffusion and structural space has already been computed as part of the previous pre-processing step (unless you ran 'dt_recon' with the --no-reg flag) and can be found in the 'register.dat' file.
The 'register.dat' file describes a rigid-body transformation from the diffusion to the structural space. To resample the structural volume into the diffusion space, we will need to apply the inverse of this transformation to our input volumes. We use the 'mri_vol2vol' command for this task. In the example below, we resample the white-matter parcellation volume (wmparc.mgz) into the diffusion space. Additionally, you may also want to repeat that process for the cortically-labelled volume (aparc+aseg.mgz).
mri_vol2vol --mov $TUTORIAL_DIR/$subj/dtrecon/lowb.nii \
--targ $SUBJECTS_DIR/$subj/mri/wmparc.mgz \
--inv --interp nearest --o $SUBJECTS_DIR/$subj/mri/wmparc2diff.mgz \
--reg $TUTORIAL_DIR/$subj/dtrecon/register.dat --no-save-regNotes:
- Since it is the inverse transformation that we apply, the role of the 'moving' and 'target' volumes are going to be inverted. The input flags should be used to reflect the original direction of the transformation, and it is the '--inv' flag that will reverse their roles.
- When computing the 'register.dat' transformation, it was the 'lowb.nii' volume that was moved into the structural space, so we indicate this volume by the '--mov' flag. (Note, any diffusion related volume from the dt_recon file could be used here.)
- The target of the original registration was in the structural space and that is indicated by the '--targ' flag. As it is the wmparc.mgz volume that we wish to resample in the diffusion space, in the above, we identify the target volume to be just that.
- The '--interp' flag indicates the type of interpolation that we want to use for the resampling. In the case of segmentation volumes, that option should always be set to nearest, indicating nearest neighbor interpolation.
- The '--o' flag indicates the desired output filename location, which in our example is 'wmparc2diff.mgz'.
- The rigid-body transformation matrix is stored in the 'register.dat' file and it is indicated by the '--reg' flag. You can use the same 'register.dat' file to move other structural volumes of this subject into the same diffusion space.
- The '--no-save-reg' flag tells 'mri_vol2vol' not to write out another registration matrix file.
To see a script which could be used to run the mri_vol2vol command on multiple subjects using both the wmparc.mgz and aparc+aseg.mgz, take a look at the AlignAnat2Diff.csh script on this page.
The above command was also run to resample the aparc+aseg for the same subject. To view the results, run:
freeview -v $TUTORIAL_DIR/$subj/dtrecon/fa.nii \
$SUBJECTS_DIR/$subj/mri/aparc+aseg2diff.mgz:colormap=lutNotes:
- The resampled aparc+aseg volume is called aparc+aseg2diff.mgz.
The command above loads the resampled aparc+aseg as well as the FA volume (fa.nii) into Freeview.
To be able to see the color labels on the aparc+aseg2diff.mgz, you must specify a color table that has those labels, colors, and names. This is done with :colormap=lut attached to the volume name that the table is needed for.
Coronal view of the cortically-labelled volume (aparc+aseg2diff.mgz) overlayed on the fractional anistropy volume (fa.nii):
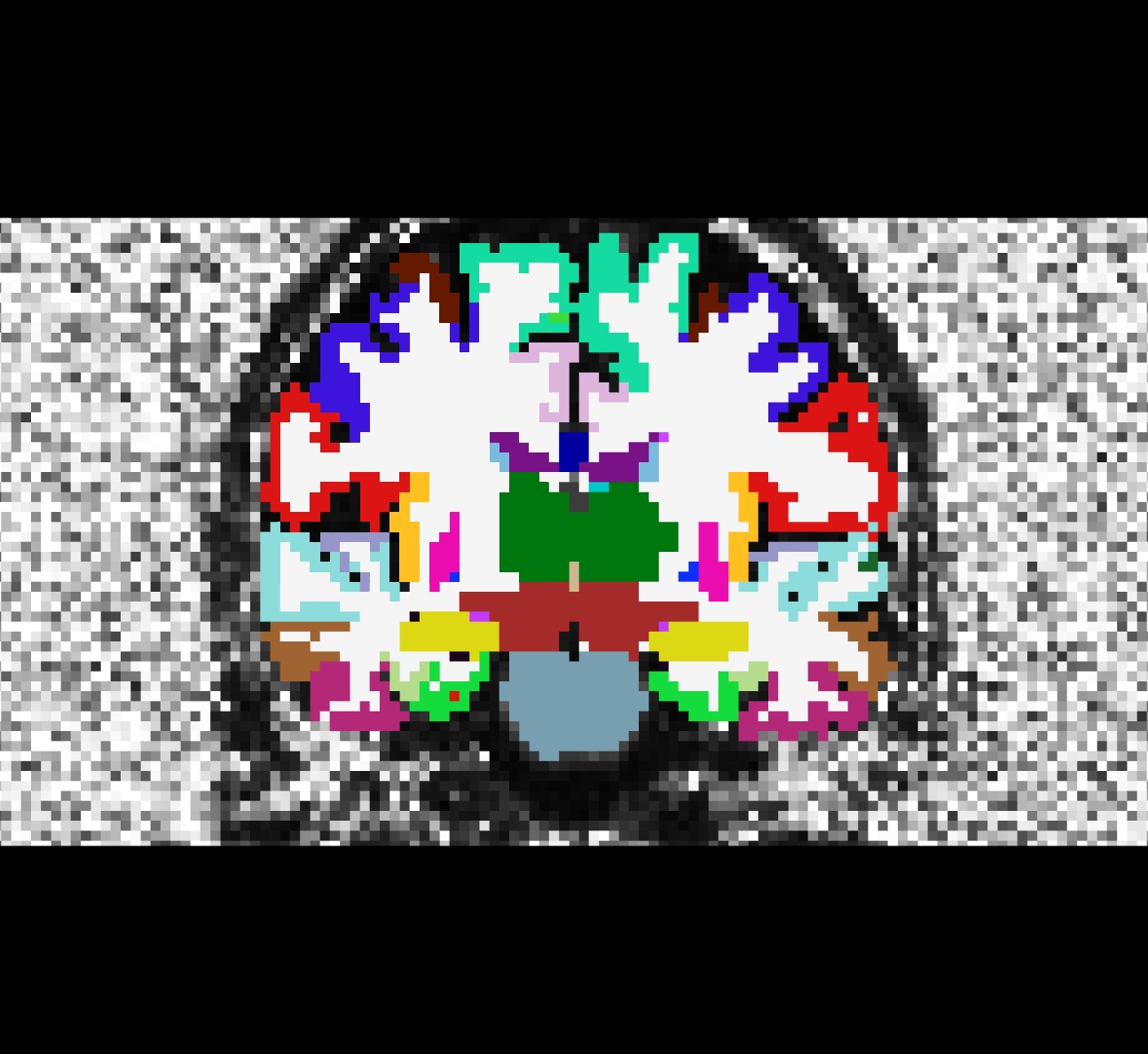

Be sure to close Freeview before proceeding to the next step.
Masking
You may have noticed that there is some unwanted "noise" or "speckles" in the diffusion images when we view them in Freeview. This can be removed by masking out the space surrounding the brain. For a mask, we will use one of the segmentation volumes from the recon directory that was already resampled into the diffusion space (wmparc2diff.mgz). To apply the mask to the diffusion data and remove the unwanted "noise", we will use the 'mri_mask' command.
mri_mask $TUTORIAL_DIR/$subj/dtrecon/fa.nii \
$SUBJECTS_DIR/$subj/mri/wmparc2diff.mgz \
$TUTORIAL_DIR/$subj/dtrecon/fa-masked.mgzNotes:
- The order of the command arguments determines the action taken. The first argument should be the volume you wish to mask. The second argument is the volume to be used as a mask. And the third argument should be the name of the output file you would like to be created. In this example, we append '-masked' to the name of the original unmasked volume file so we can tell them apart.
- The command above applies a mask to the fractional anisotropy volume (fa.nii), but you may run 'mri_mask' on any other diffusion volume (i.e., adc.nii, ivc.nii)
- All non-brain areas will be assigned to the background.
To see a script which could be used to run the mri_mask command on multiple subjects take a look at the 'DiffMasking.csh' script on this page.
Run the freeview command below to compare the fractional anisotropy volume (fa.nii) before and after masking:
freeview -v $TUTORIAL_DIR/$subj/dtrecon/fa.nii \
$TUTORIAL_DIR/$subj/dtrecon/fa-masked.mgzUse Alt+c to quickly alternate between the two volumes and see the difference.
Coronal view of the unmasked fractional anisotropy volume (fa.nii):
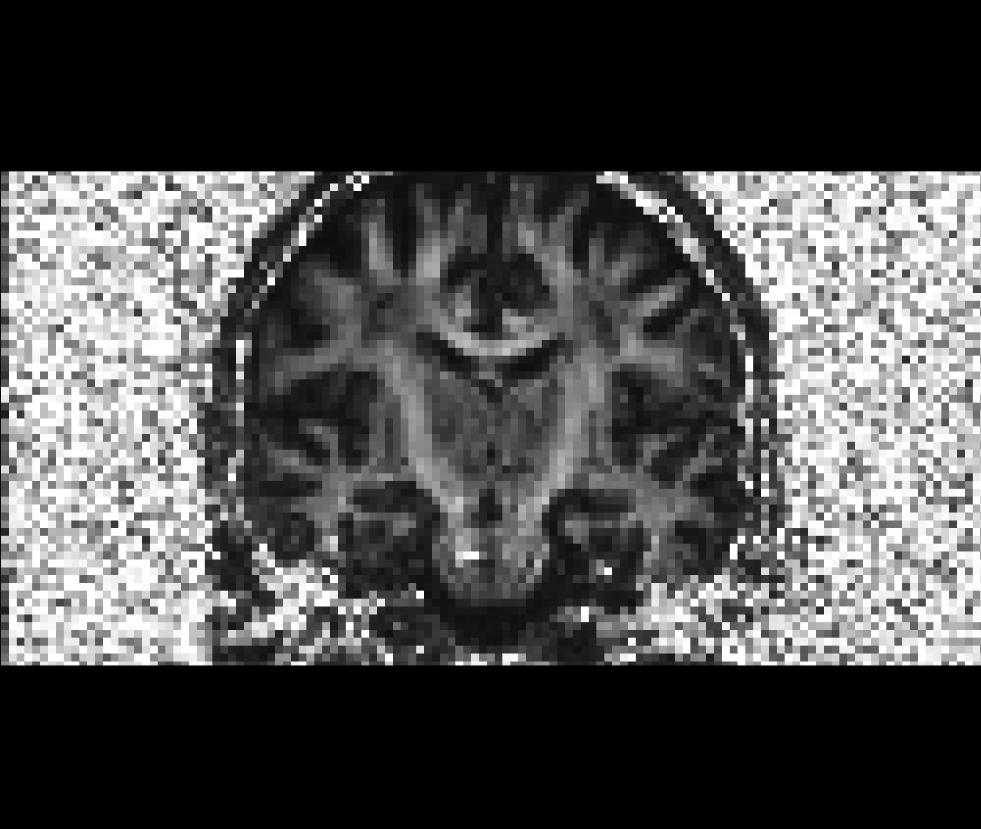
Coronal view of the masked fractional anisotropy volume (fa-masked.mgz):
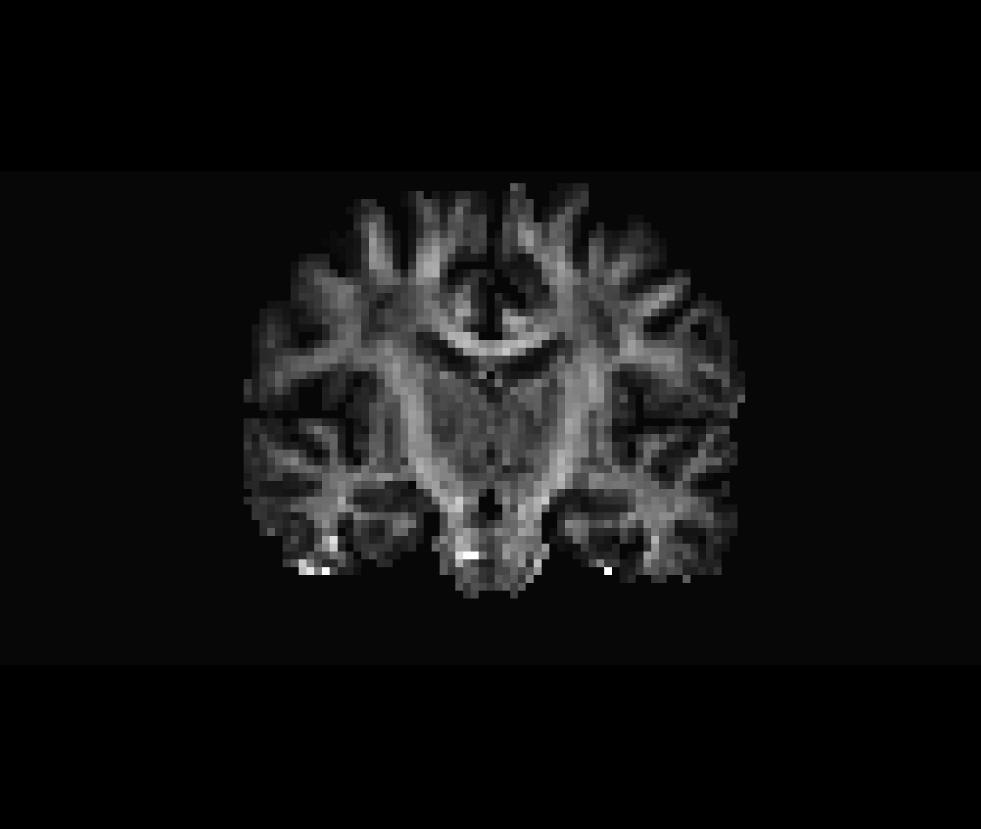
Be sure to close the Freeview window before proceeding with the next step.
Spatial Normalization
At this point we could run analyses on a subject's DTI and structural data, but we could not do a voxel-based group analysis with multiple subjects, because the data sets are not yet aligned in a common coordinate space. In other words, a certain coordinate location in one subject's volume does not necessarily correspond to the same anatomical location in any of our other subjects. We want to normalize (resample) the diffusion data of each subject into a specified common coordinate space.
In the example below, the data will be resampled into CVS space, but alternatively you could normalize your data in any other coordinate system (for example, MNI or Talairach). As a result of running mri_cvs_register on the structural acquisitions, a .m3z morph file ('combined_tocvs_avg35_elreg_afteraseg-norm.m3z') is created which describes the transformation for each subject that is necessary to move from an individual's native space to that of the common CVS space.
!!! WARNING: DO NOT RUN THIS COMMAND ON THE COURSE LAPTOPS !!!
This command currently requires greater than 4GB of memory to run. We have already run it for you.
mri_vol2vol --targ $FREESURFER_HOME/subjects/cvs_avg35/mri/norm.mgz \
--m3z $SUBJECTS_DIR/$subj/cvs/combined_tocvs_avg35_elreg_afteraseg-norm.m3z \
--noDefM3zPath --reg $TUTORIAL_DIR/$subj/dtrecon/register.dat \
--mov $TUTORIAL_DIR/$subj/dtrecon/fa-masked.mgz \
--o $TUTORIAL_DIR/$subj/dtrecon/fa-masked.ANAT+CVS-to-avg35.mgz \
--interp trilin --no-save-regNotes:
- The '--mov' flag indicates the moving volume, the volume we want to resample in the common CVS space for a subsequent group analysis.
- The target CVS space where we want to resample our moving volume (fa-masked.mgz) is indicated with the '--targ' flag.
- The '--m3z' flag indicates the name of a morph file created for this subject by the mri_cvs_register command, which serves as a link between this subject's native space and the CVS space.
- The '--noDefM3zPath' flag indicates to mri_vol2vol not to search for the morph in a default directory ($SUBJECTS_DIR/$subj/mri/transforms) but to instead use the full input filename.
- The '--reg' flag identifies the rigid-body motion that describes the connection between the structural and the diffusion spaces. As the CVS morph is computed using structural volumes and in the above example we are applying the CVS morph to a diffusion volume, we need to make sure that the transformation between these two spaces is also taken into account during the resampling process.
- The '--o' flag describes the desired name of the output file.
- The '--interp' flag indicates the type of interpolation we want to use for our resampling. We are not using a segmented volume as an input so we use the default trilinear (trilin) interpolation option. If you wanted to resample a segmentation volume (such as wmparc or aseg) in CVS space, you would want to use the nearest neighbor (nearest) interpolation option.
- Again, the '--no-save-reg' flag tells the command not to write out another copy of the registration matrix that is being applied.
You would need to run this command on every subject in your data set before running a group analysis. To see a script which could be used to do that, take a look at the !AlignAnatCVSToAVG.csh script on this page.
To view the 'norm.mgz' file (which is in the target CVS space) and the 'fa-masked.mgz' file after is has been resampled in the common CVS space, run:
freeview -v $FREESURFER_HOME/subjects/cvs_avg35/mri/norm.mgz \
$TUTORIAL_DIR/$subj/dtrecon/fa-masked.ANAT+CVS-to-avg35.mgzUse Alt+c to alternate between the two volumes.
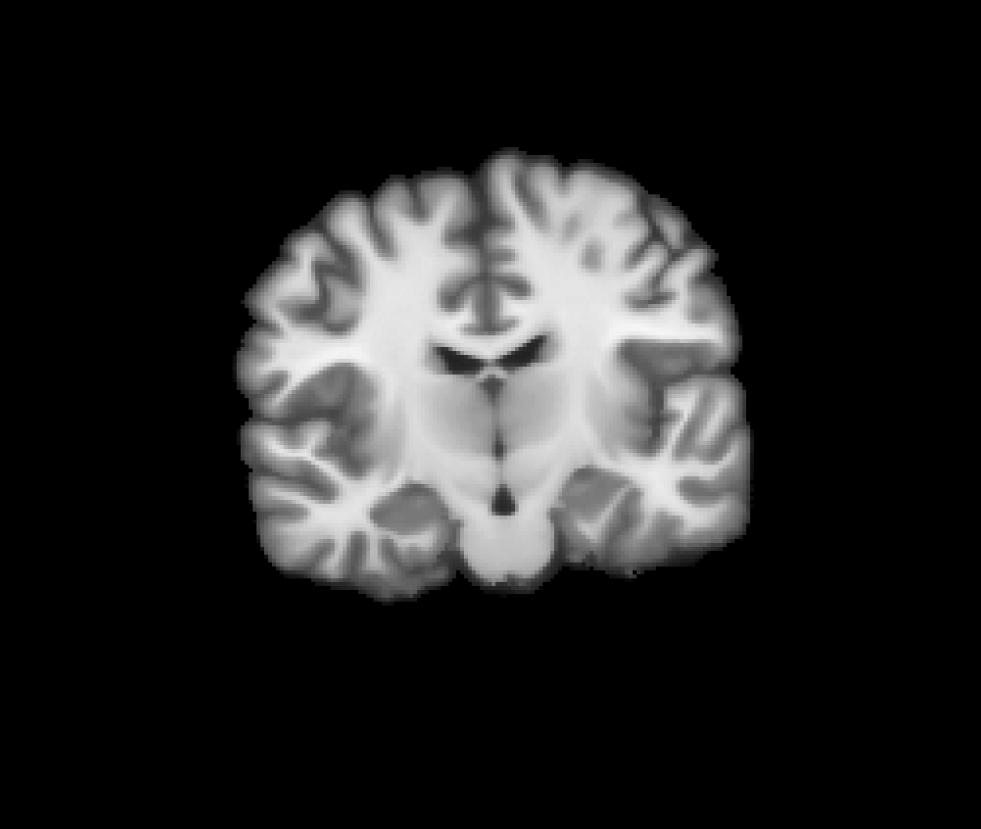
Coronal view of the target CVS space (norm.mgz) we resampled into:
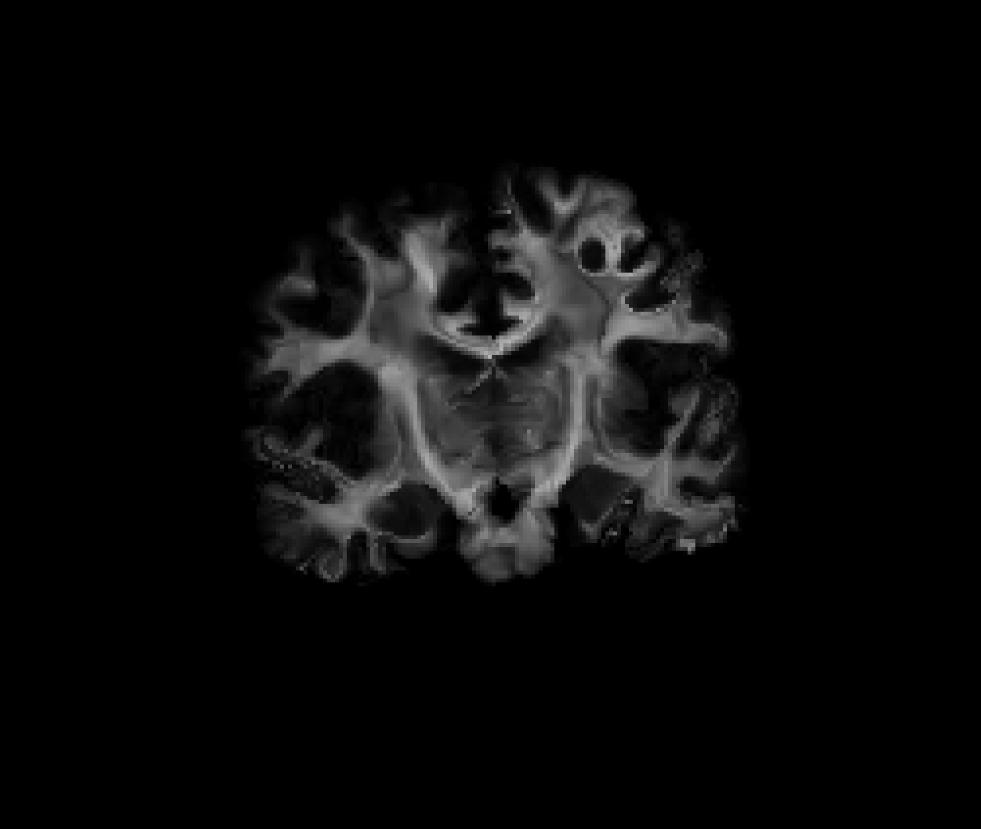
Coronal view of the masked fractional anisotropy volume after it was resampled int CVS space (fa-masked.ANAT+CVS-to-avg35.mgz):
Once all the subjects in your study are put into the same coordinate system, you are ready to begin your group analysis on the resampled subjects.
Links Of Interest
Acknowledgements
DTI tutorial scripts created by Lilla Zollei.
