SynthStrip: Skull-Stripping for Any Brain Image
Andrew Hoopes, Jocelyn S. Mora, Adrian V. Dalca, Bruce Fischl, Malte Hoffmann
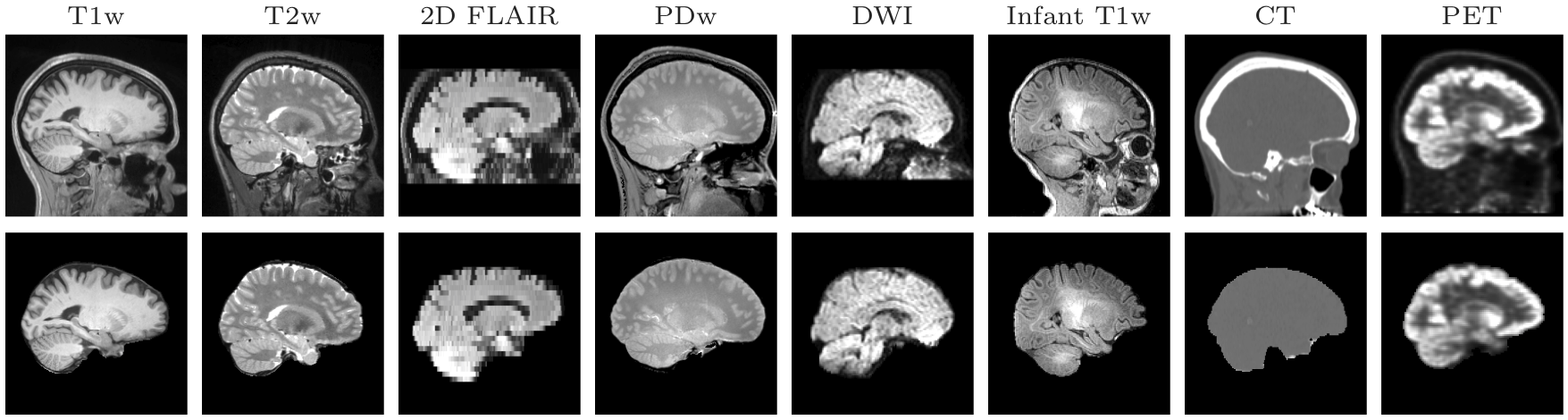
SynthStrip is a skull-stripping tool that extracts brain voxels from a landscape of image types, ranging across imaging modalities, resolutions, and subject populations. It leverages a deep learning strategy to synthesize arbitrary training images from segmentation maps, yielding a robust model agnostic to acquisition specifics.
Publications
If you find this work useful, please cite the relevant papers below (BibTeX).
Main method, toolbox, and dataset:
SynthStrip: Skull-Stripping for Any Brain Image
Andrew Hoopes, Jocelyn S. Mora, Adrian V. Dalca, Bruce Fischl*, Malte Hoffmann* (*equal contribution)
NeuroImage, 260, p 119474, 2022
Pediatric model:
Boosting Skull-Stripping Performance for Pediatric Brain Images
William Kelley, Nathan Ngo, Adrian V. Dalca, Bruce Fischl, Lilla Zöllei*, Malte Hoffmann* (*equal contribution)
IEEE International Symposium on Biomedical Imaging (ISBI), pp 1-5, 2024
Synthesis-driven training and domain randomization:
Domain-Randomized Deep Learning for Neuroimage Analysis
Malte Hoffmann
IEEE Signal Processing Magazine (SPM), 42 (4), pp 78-90, 2025

SynthStrip Dataset
The SynthStrip dataset is a collection of full-head images with associated ground-truth brain masks from 622 MRI, CT, and PET scans. We include label maps for 131 adult MPRAGE scans, with standard FreeSurfer brain labels and additional non-brain labels. The images span various MRI contrasts, resolutions, and populations ranging from infants to glioblastoma patients. While we cannot redistribute the CT and PET data, we provide information on how to obtain these. The 2D subset consists of sagittal slices extracted from each file in the 3D dataset.
See README for dataset information and license
Download the 3D SynthStrip dataset (v1.5, 6.9 GB)
Download the 2D SynthStrip dataset (v1.5, 39 MB)
Verify download integrity with SHA-256 checksums
Download, verify, and extract the data with:
curl -O https://surfer.nmr.mgh.harvard.edu/docs/synthstrip/data/SHA256
curl -O https://surfer.nmr.mgh.harvard.edu/docs/synthstrip/data/synthstrip_data_v1.5.tar
curl -O https://surfer.nmr.mgh.harvard.edu/docs/synthstrip/data/synthstrip_data_v1.5_2d.tar
shasum -c SHA256
tar -xf synthstrip_data_v1.5.tar
tar -xf synthstrip_data_v1.5_2d.tarIf you use these data, please cite SynthStrip (BibTeX).
SynthStrip Tool
We ship SynthStrip as a command-line tool with FreeSurfer and as a standalone utility using Docker or Singularity containers. Both versions are functionally identical and use the same command-line syntax.
Within FreeSurfer: The mri_synthstrip utility has been included in FreeSurfer since the v7.3.0 release. For the most up-to-date version of SynthStrip, please download a build of the FreeSurfer development branch.
Container image: If you do not want to install FreeSurfer, you can run SynthStrip in a container. We provide a wrapper script, so you do not need to mount input and output directories. The image is available on Docker Hub.
Apptainer or Singularity: Download the Singularity-based wrapper script with:
curl -O https://raw.githubusercontent.com/freesurfer/freesurfer/dev/mri_synthstrip/synthstrip-singularity && chmod +x synthstrip-singularityDocker: Download the Docker-based wrapper script with:
curl -O https://raw.githubusercontent.com/freesurfer/freesurfer/dev/mri_synthstrip/synthstrip-docker && chmod +x synthstrip-dockerPlease read the instructions at the top of the downloaded script. Singularity requires simple one-time configuration.
Usage
Once installed, run SynthStrip as follows, where "stripped.nii.gz" is a skull-stripped version of the image "input.nii.gz".
mri_synthstrip -i input.nii.gz -o stripped.nii.gzNote: For the container version, replace mri_synthstrip with the wrapper script name (e.g. synthstrip-singularity).
Use the -m flag to save a binary brain mask:
mri_synthstrip -i input.mgz -o stripped.mgz -m mask.mgzIf you would like to compute the immediate boundary of the brain excluding surrounding CSF, use the --no-csf flag. You may also want to explore the -b option, which controls the boundary distance from the brain.
mri_synthstrip -i input.nii -o stripped.nii --no-csfDisplay additional options with the --help flag. SynthStrip should take less than 1 minute on the CPU for most images with voxel sizes near 1 mm3. As image size or resolution increases, the runtime might increase as well.
Video (5 minutes)
Code and Weights
For a custom Python setup, download the SynthStrip script from FreeSurfer's GitHub repository. You can choose to access the weight files under either the MIT license or the CC BY 4.0 license:
- the main SynthStrip model (version 1, 29 MB)
- a model for predicting brain masks without CSF (version 1, 29 MB)
- a pediatric SynthStrip model for brain masks without CSF (version 1, 29 MB)
We also export the Python requirements used to build the latest SynthStrip container.
Changes
A list of changes and bug fixes is available on Docker Hub. For ease of use, download a recent FreeSurfer version or update the top of the wrapper script when using the container image.
Acknowledgments
The authors thank Douglas Greve and David Salat for sharing data. This research project benefitted from computational hardware generously provided by the Massachusetts Life Sciences Center.