Steps for Using the Diffusion Dialog Table
note: All the source code can be downloaded from the "dev" branch on the free surfer githhub page. Refer to this link to explore the code used for this tool: https://github.com/freesurfer/freesurfer/tree/dev/diffusion_tool.
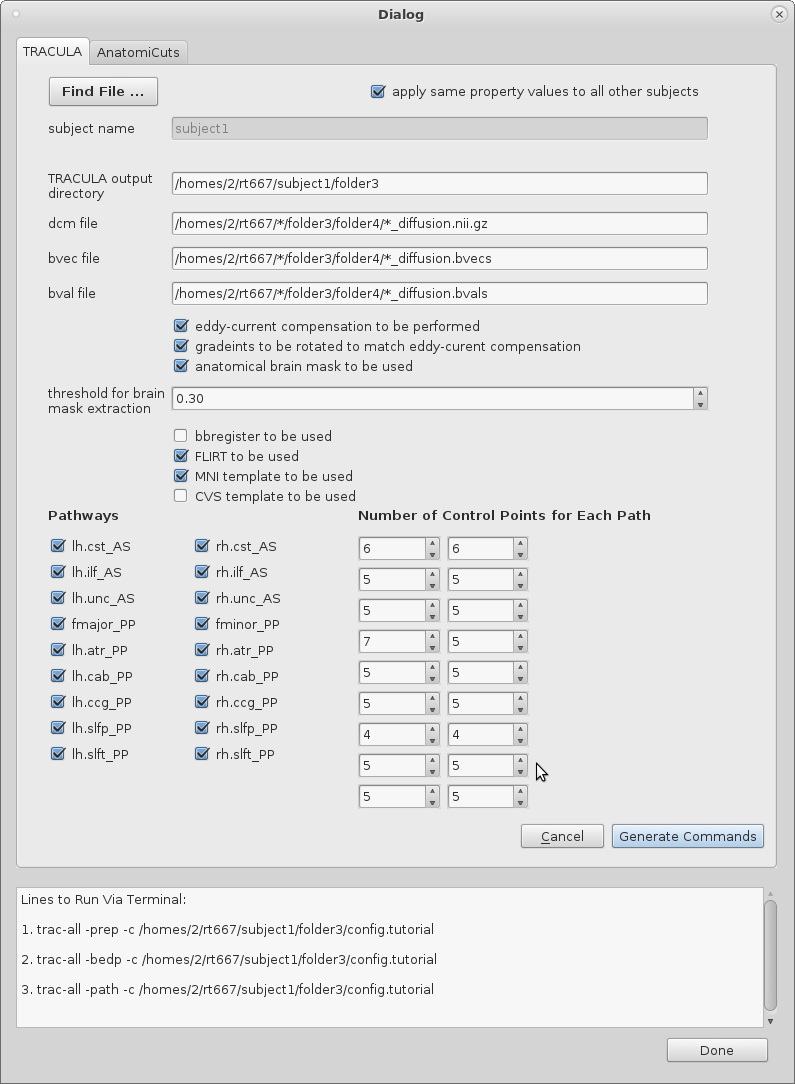
Running TRACULA:
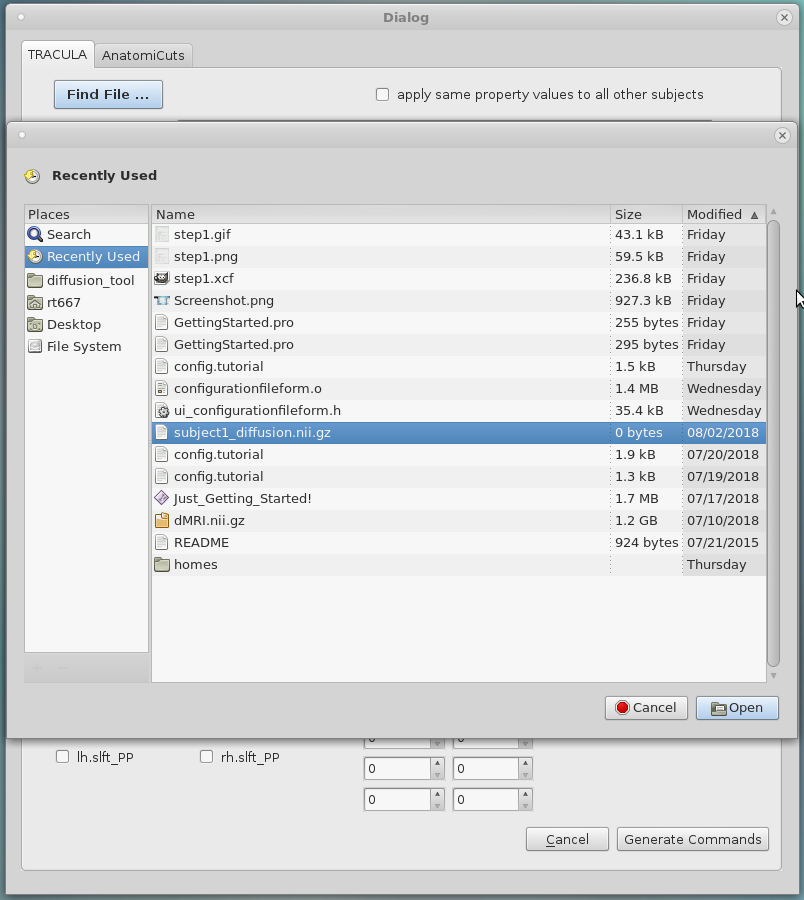
1. Select the "Find File..." button to open another window where a DCOM file with the appropriate extension must be selected. If the file does not have either a ".nii.gz" or ".dcm" extension, a warning message will be outputted and the file will not be accepted into the dialog table.
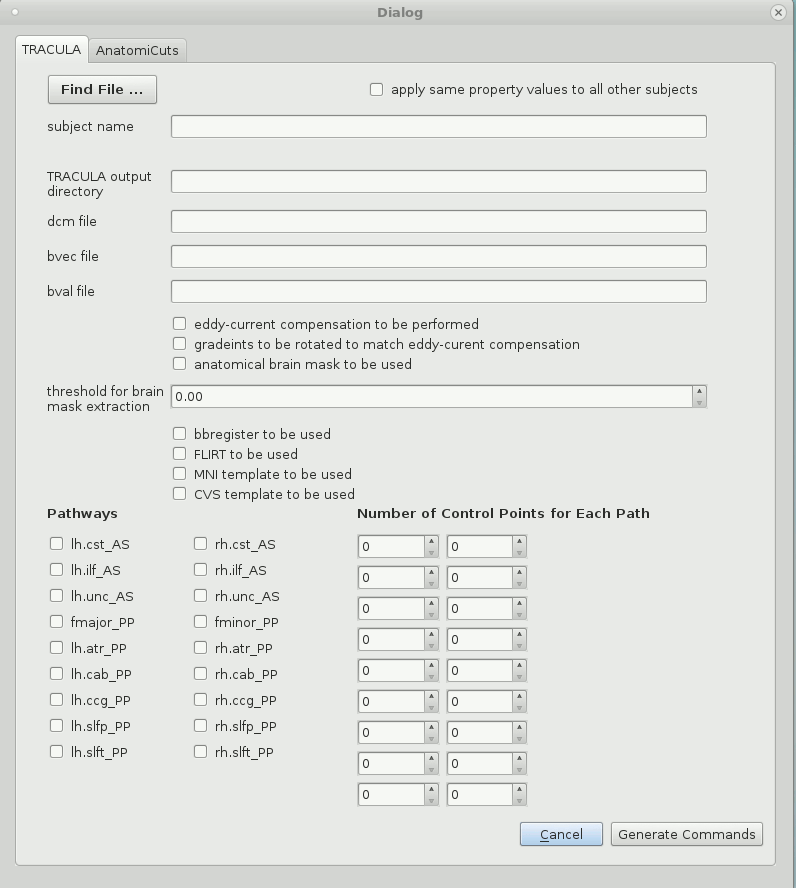
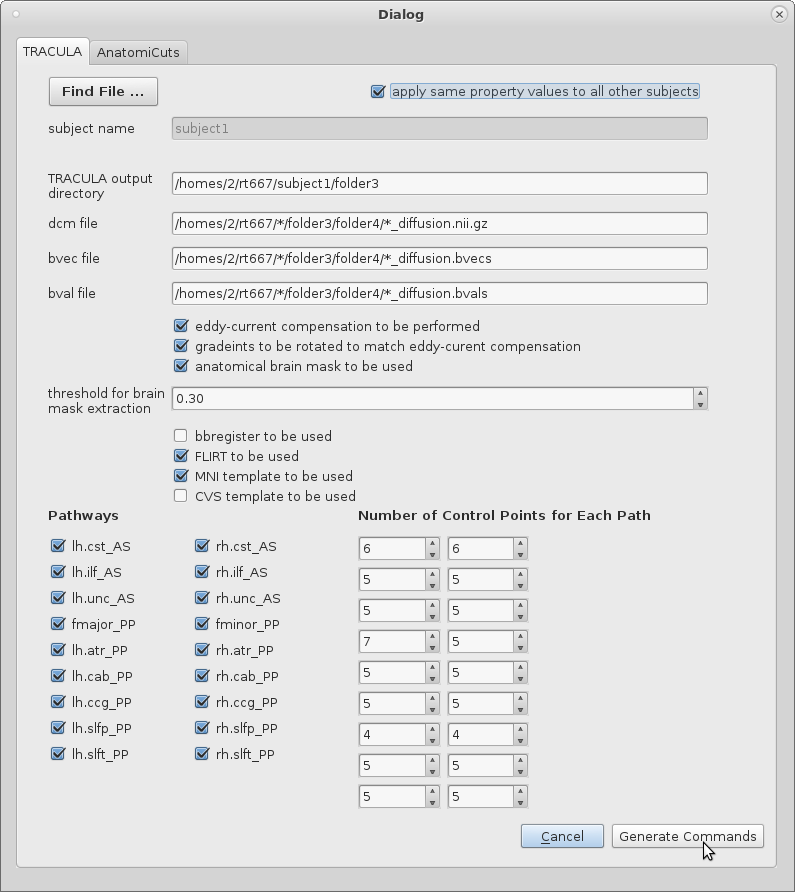
2. A default subject name will have been chosen by the program. Correct and edit the string that represents the subject name if necessary.
3. Check the box with the label "apply same property values to all other subjects" if TRACULA is expected to be run on a number of subjects who all share a pattern in the strings that represent paths to their DCOM, bval, and bvec files.
4. String will have been loaded into the dialog table that will represent the path to the DCOM, bval, and bvec files. Additionally, a string representing the path to the "output directory" where the configuration file will be written to under the name "config.tutorial" will be written. If these defaults are to be changed, they must be done so by the user. If the user had selected the check box that applies the same property values to all other subjects, any instance within the strings where the subject's name would have appeared will have been replaced with a star to confirm generality.
5. The remaining check boxes and input values will have been set to certain defaults that can be changed easily by the user. These are all variables that will be necessary to include and specify in the configuration file, and thereby the running of TRACULA itself.
6. Once the table is complete with appropriate values, the user can select the "Generate Command" to both write the configuration file (which will be written into the output directory) and display the three lines that must be run at the command-line on terminal. These commands will handle the pre-processing of the diffusion image data as well as the reconstructing of the white-matter pathways. Run these commands to run TRACULA.
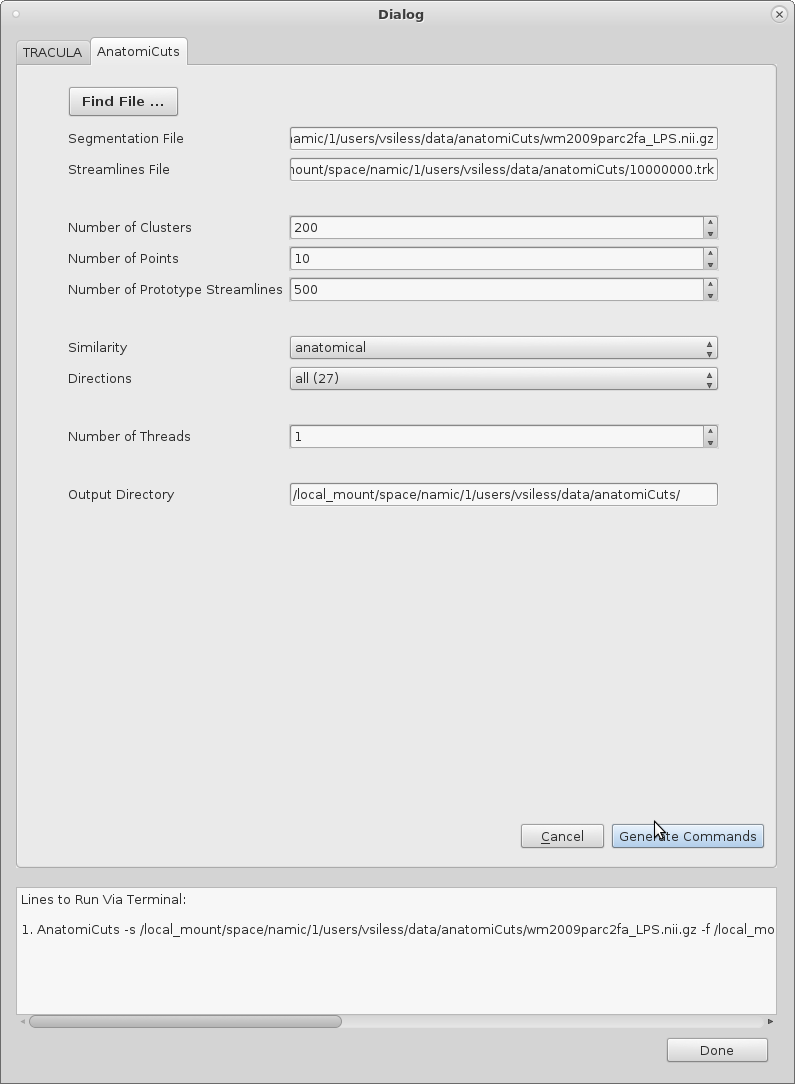
Running AnatomiCuts
1. Select the "Find File..." button to open another window where a nifty file with the appropriate extension must be selected. If the file does not have a ".nii.gz", a warning message will be outputted and the file will not be accepted into the dialog table.
2. Upon selecting a nifty file, a streamlines file with a ".trk" extension will be searched for. If it is found, the rest of the options in the dialog table will be set to default values. These strings and values can always be changed by the user.
3. Once the entire table's values have been set, the user can select the button "Generate Commands" and a string representing the command to be run on terminal will be displayed. Run this command to run AnatomiCuts.