|
Size: 15735
Comment:
|
Size: 15602
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| Line 29: | Line 30: |
This tutorial is designed to introduce you to the "command-line" group analysis stream in FreeSurfer (as opposed to QDEC which is GUI-driven), including correction for multiple comparisons. While this tutorial shows you how to perform a surface-based thickness study, it is important to realize that most of the concepts learned here apply to any group analysis in FreeSurfer, surface or volume, thickness or fMRI. Here are some useful Group Analysis Links: <<BR>> [[FsTutorial/QdecGroupAnalysis|QDEC Tutorial]]<<BR>> [[FsgdFormat | FSGD Format]]<<BR>> [[FsgdExamples|FSGD Examples]]<<BR>> [[DodsDoss| DODS vs DOSS]]<<BR>> |
This tutorial is designed to introduce you to the "command-line" group analysis stream in FreeSurfer (as opposed to QDEC which is GUI-driven), including correction for multiple comparisons. While this tutorial shows you how to perform a surface-based thickness study, it is important to realize that most of the concepts learned here apply to any group analysis in FreeSurfer, surface or volume, thickness or fMRI. Here are some useful Group Analysis Links: <<BR>> [[FsTutorial/QdecGroupAnalysis|QDEC Tutorial]]<<BR>> [[FsgdFormat|FSGD Format]]<<BR>> [[FsgdExamples|FSGD Examples]]<<BR>> [[DodsDoss|DODS vs DOSS]]<<BR>> |
| Line 44: | Line 35: |
| Line 46: | Line 36: |
The data used for this tutorial is 40 subjects from Randy Buckner's lab. It consists of males and females ages 18 to 93. You can see the demographics [[FsTutorial/GroupAnalysisDemographics|here]]. You will perform an analysis looking for the effect of age on cortical thickness accounting for the effects of gender in the analysis. This is the same data set used in the [[FsTutorial/QdecGroupAnalysis|QDEC Tutorial]], and you will get the same result. |
The data used for this tutorial is 40 subjects from Randy Buckner's lab. It consists of males and females ages 18 to 93. You can see the demographics [[FsTutorial/GroupAnalysisDemographics|here]]. You will perform an analysis looking for the effect of age on cortical thickness accounting for the effects of gender in the analysis. This is the same data set used in the [[FsTutorial/QdecGroupAnalysis|QDEC Tutorial]], and you will get the same result. |
| Line 57: | Line 39: |
| Line 59: | Line 40: |
For this example, we will model the thickness as a straight line. A line has two parameters: intercept (or offset) and a slope. 1. The slope is the change of thickness with age 1. The intercept/offset is interpreted as the thickness at age=0. |
For this example, we will model the thickness as a straight line. A line has two parameters: intercept (or offset) and a slope. 1. The slope is the change of thickness with age 1. The intercept/offset is interpreted as the thickness at age=0. |
| Line 66: | Line 47: |
| To account for effects of gender, we will model each sex with its own line, meaning that there will be four linear parameters (also called "betas"): 1. Intercept for Females 1. Intercept for Males 1. Slope for Females 1. Slope for Males In FreeSurfer, this type of design is called [[DodsDoss| DODS]] (for "Different-Offset, Different-Slope"). In FreeSurfer, you can either create your own design matrices, or, if you can specify your design in terms of a FreeSurfer Group Descriptor File [[FsgdFormat| (FSGD)]], FreeSurfer will create them for you. The FSGD file is a simple text file. See [[FsgdFormat|this page]] for the format. The [[FsTutorial/GroupAnalysisDemographics|demographics page]] also has an example FSGD file for this data. |
To account for effects of gender, we will model each sex with its own line, meaning that there will be four linear parameters (also called "betas"): 1. Intercept for Females 1. Intercept for Males 1. Slope for Females 1. Slope for Males In FreeSurfer, this type of design is called [[DodsDoss|DODS]] (for "Different-Offset, Different-Slope"). In FreeSurfer, you can either create your own design matrices, or, if you can specify your design in terms of a FreeSurfer Group Descriptor File [[FsgdFormat|(FSGD)]], FreeSurfer will create them for you. The FSGD file is a simple text file. See [[FsgdFormat|this page]] for the format. The [[FsTutorial/GroupAnalysisDemographics|demographics page]] also has an example FSGD file for this data. |
| Line 84: | Line 59: |
| 1. Create an FSGD file for the above design. One (gender_age.fsgd) already exists so that you can continue with the exercises. Information on how to create/view text files can be found [[FsTutorial/CreatingTextfiles|here]]. |
1. Create an FSGD file for the above design. One (gender_age.fsgd) already exists so that you can continue with the exercises. Information on how to create/view text files can be found [[FsTutorial/CreatingTextfiles|here]]. |
| Line 90: | Line 65: |
A contrast is a vector that embodies the hypothesis we want to test. In this case, we wish to test the change in thickness with age after removing the effects of gender. To do this, create a simple text file with the following numbers: |
A contrast is a vector that embodies the hypothesis we want to test. In this case, we wish to test the change in thickness with age after removing the effects of gender. To do this, create a simple text file with the following numbers: |
| Line 99: | Line 70: |
Notes: 1. There is one value for each parameter (so 4 values) 1. The intercept/offset values are 0 (nuisance) 1. The slope values are 0.5 so as to average the F and M slopes 1. A file called lh-Avg-thickness-age-Cor.mtx already exists |
Notes: 1. There is one value for each parameter (so 4 values) 1. The intercept/offset values are 0 (nuisance) 1. The slope values are 0.5 so as to average the F and M slopes 1. A file called lh-Avg-thickness-age-Cor.mtx already exists |
| Line 107: | Line 78: |
| Line 109: | Line 79: |
| Line 111: | Line 80: |
| 1. Resampling each subject's data into a common space, and 1. Concatenating all the subjects' into a single file 1. Spatial smoothing (can be done between 1 and 2) |
1. Resampling each subject's data into a common space, and 1. Concatenating all the subjects' into a single file 1. Spatial smoothing (can be done between 1 and 2) |
| Line 117: | Line 88: |
When you have run recon-all with the -qcache option, recon-all will resample data onto the average subject (fsaverage) and smooth it at various FWHM (full-width/half-max), usually 0, 5, 10, 10, 20, and 25mm. This can speed later processing. The data for this tutorial have been cached, so run: |
When you have run recon-all with the -qcache option, recon-all will resample data onto the average subject (fsaverage) and smooth it at various FWHM (full-width/half-max), usually 0, 5, 10, 10, 20, and 25mm. This can speed later processing. The data for this tutorial have been cached, so run: |
| Line 129: | Line 96: |
}}} ---- Notes: 1. Only takes a few seconds because the data have been cached 1. The FSGD file lists all the subjects, helps keep order. 1. The independent variable is the thickness smoothed to 10mm FWHM. 1. The data are for the left hemisphere 1. The output is lh.gender_age.thickness.10.mgh (which already exists) |
}}} ---- Notes: 1. Only takes a few seconds because the data have been cached 1. The FSGD file lists all the subjects, helps keep order. 1. The independent variable is the thickness smoothed to 10mm FWHM. 1. The data are for the left hemisphere 1. The output is lh.gender_age.thickness.10.mgh (which already exists) |
| Line 141: | Line 107: |
| 1. Run mri_info on lh.gender_age.thickness.10.mgh to find its dimensions,ie, ---- |
1. Run mri_info on lh.gender_age.thickness.10.mgh to find its dimensions,ie, ---- |
| Line 149: | Line 116: |
In the case that you have not cached the data, you can perform the same operations manually using the two commands below. There is no need to do this for this tutorial. OPTIONAL: THIS WILL TAKE ABOUT 10 MINUTES |
In the case that you have not cached the data, you can perform the same operations manually using the two commands below. There is no need to do this for this tutorial. OPTIONAL: THIS WILL TAKE ABOUT 10 MINUTES |
| Line 161: | Line 124: |
Notes: 1. This resamples each subjects data to fsaverage 1. Output is lh.gender_age.thickness.00.mgh, which is unsmoothed |
Notes: 1. This resamples each subjects data to fsaverage 1. Output is lh.gender_age.thickness.00.mgh, which is unsmoothed |
| Line 177: | Line 139: |
1. This smooths by 10mm FWHM 1. "--cortex" means only smooth areas in cortex (exclude medial wall). This is automatically done with qcache. You can also specify other labels. 1. Output is lh.gender_age.thickness.10B.mgh. |
1. This smooths by 10mm FWHM 1. "--cortex" means only smooth areas in cortex (exclude medial wall). This is automatically done with qcache. You can also specify other labels. 1. Output is lh.gender_age.thickness.10B.mgh. |
| Line 185: | Line 144: |
| Line 195: | Line 153: |
}}} ---- Notes: 1. Input is lh.gender_age.thickness.10.mgh 1. Same FSGD used as with mris_preproc. Maintains subject order! 1. DODS is specified (it is the default) 1. Only one contrast is used (lh-Avg-thickness-age-Cor.mtx) but you can specify multiple contrasts. 1. "--cortex" specifies that the analysis only be done in cortex (ie, medial wall is zeroed out). Other labels can be used. 1. The output directory is lh.gender_age.glmdir 1. Should only take about 1min to run. |
}}} ---- Notes: 1. Input is lh.gender_age.thickness.10.mgh 1. Same FSGD used as with mris_preproc. Maintains subject order! 1. DODS is specified (it is the default) 1. Only one contrast is used (lh-Avg-thickness-age-Cor.mtx) but you can specify multiple contrasts. 1. "--cortex" specifies that the analysis only be done in cortex (ie, medial wall is zeroed out). Other labels can be used. 1. The output directory is lh.gender_age.glmdir 1. Should only take about 1min to run. |
| Line 211: | Line 167: |
| When this command is finished the '''glm''' directory will contain a sub-directory called '''lh.gender_age.glmdir'''. There will be a number of output files in this directory, as well as two other directories. If you did an ''ls'' in your glmdir there will be: |
When this command is finished the '''glm''' directory will contain a sub-directory called '''lh.gender_age.glmdir'''. There will be a number of output files in this directory, as well as two other directories. If you did an ''ls'' in your glmdir there will be: |
| Line 228: | Line 181: |
| Line 230: | Line 182: |
| 1. What was the DOF for this experiment? 1. What was the FWHM? 1. How many frames does beta have? Why? There will be a subdirectory for each contrast that you specify. The name of the directory will be that of the contrast matrix file (without the .mtx extension). The '''lh-Avg-thickness-age-Cor''' directory will have the following files: |
1. What was the DOF for this experiment? 1. What was the FWHM? 1. How many frames does beta have? Why? There will be a subdirectory for each contrast that you specify. The name of the directory will be that of the contrast matrix file (without the .mtx extension). The '''lh-Avg-thickness-age-Cor''' directory will have the following files: |
| Line 245: | Line 195: |
| Line 247: | Line 196: |
| Line 252: | Line 202: |
| Line 259: | Line 208: |
| {{attachment:glm_1.jpg}} {{attachment:glm_2.jpg}} Notes: 1. The threshold is set to 2, meaning vertices with p<.01, uncorrected, will have color. 1. Blue means a negative correlation (thickness decreases with age), red is positive correlation. 1. Click on a point in the Precentral Gyrus. What is it's value? What does it mean? Viewing the medial surface, change the overlay threshold to something very, very low (say, .01), by clicking '''Configure Overlay''' and set the '''Min''' value to 0.01. Click Apply, or check the '''Automatically apply changes''' checkbox. |
{{attachment:glm1_fv.jpg||height="370",width="480"}} {{attachment:glm2_fv.jpg||height="370",width="553"}} Notes: 1. The threshold is set to 2, meaning vertices with p<.01, uncorrected, will have color. 1. Blue means a negative correlation (thickness decreases with age), red is positive correlation. 1. Click on a point in the Precentral Gyrus. What is it's value? What does it mean? Viewing the medial surface, change the overlay threshold to something very, very low (say, .01), by clicking '''Configure Overlay''' and set the '''Min''' value to 0.01. Click Apply, or check the '''Automatically apply changes''' checkbox. |
| Line 275: | Line 220: |
| {{attachment:glm_3.jpg}} Notes: 1. Almost all of cortex now has color 1. The non-cortical areas are still blank (0) because they were excluded with --cortex in mri_glmfit above. |
{{attachment:glm3_fv.jpg}} Notes: 1. Almost all of cortex now has color 1. The non-cortical areas are still blank (0) because they were excluded with --cortex in mri_glmfit above. |
| Line 283: | Line 228: |
| Line 286: | Line 230: |
| To perform a cluster-wise correction for multiple comparisons, we will run a simulation. The simulation is a way to get a measure of the distribution of the maximum cluster size under the null hypothesis. This is done by iterating over the following steps: 1. Synthesize a z map 1. Smooth z map (using residual FWHM) 1. Threshold z map (level and sign) 1. Find clusters in thresholded map 1. Record area of maximum cluster 1. Repeat over desired number of iterations (usually > 5000) In FreeSurfer, this information is stored in a simple text file called a CSD (Cluster Simulation Data). Once we have the distribution of the maximum cluster size, we correct for multiple comparisons by: 1. Going back to the original data 1. Thresholding using same level and sign 1. Finding clusters in thresholded map 1. For each cluster, p = probability of seeing a maximum cluster that size or larger during simulation. |
To perform a cluster-wise correction for multiple comparisons, we will run a simulation. The simulation is a way to get a measure of the distribution of the maximum cluster size under the null hypothesis. This is done by iterating over the following steps: 1. Synthesize a z map 1. Smooth z map (using residual FWHM) 1. Threshold z map (level and sign) 1. Find clusters in thresholded map 1. Record area of maximum cluster 1. Repeat over desired number of iterations (usually > 5000) In FreeSurfer, this information is stored in a simple text file called a CSD (Cluster Simulation Data). Once we have the distribution of the maximum cluster size, we correct for multiple comparisons by: 1. Going back to the original data 1. Thresholding using same level and sign 1. Finding clusters in thresholded map 1. For each cluster, p = probability of seeing a maximum cluster that size or larger during simulation. |
| Line 309: | Line 249: |
| Line 321: | Line 260: |
Notes: 1. Specify the same GLM directory (--glmdir) 1. The simulation type is Z Monte Carlo (mc-z) 1. Specify the sign (neg for negative, pos, or abs) 1. Vertex-wise threshold of 4 (p < .0001) 1. Only 5 iterations (usually > 5000) 1. CSD files will have mc-z.negative base 1. "--cwpvalthresh 0.4" Keep clusters that have cluster-wise p-values < 0.4 (usually you will want something like .05). To see all clusters, set to .999 1. --overwrite deletes previous CSD files This has been run this data with 1000 iterations (CSD base of mc-z.neg). This can often take many hours. |
Notes: 1. Specify the same GLM directory (--glmdir) 1. The simulation type is Z Monte Carlo (mc-z) 1. Specify the sign (neg for negative, pos, or abs) 1. Vertex-wise threshold of 4 (p < .0001) 1. Only 5 iterations (usually > 5000) 1. CSD files will have mc-z.negative base 1. "--cwpvalthresh 0.4" Keep clusters that have cluster-wise p-values < 0.4 (usually you will want something like .05). To see all clusters, set to .999 1. --overwrite deletes previous CSD files This has been run this data with 1000 iterations (CSD base of mc-z.neg). This can often take many hours. |
| Line 336: | Line 274: |
The CSD files are stored in lh.gender_age.glmdir/csd. Look at one of them: |
The CSD files are stored in lh.gender_age.glmdir/csd. Look at one of them: |
| Line 349: | Line 286: |
| 1. Rows that begin with '#' are headers or comments. 1. Each data row is an iteration. 1. The 3rd column is the maximum cluster size for that iteration. 1. The CSD file itself is not particularly important, so don't worry too much about it. |
1. Rows that begin with '#' are headers or comments. 1. Each data row is an iteration. 1. The 3rd column is the maximum cluster size for that iteration. 1. The CSD file itself is not particularly important, so don't worry too much about it. |
| Line 356: | Line 293: |
In the contrast subdirectory, you will see several new files, all beginning with mc-z: |
In the contrast subdirectory, you will see several new files, all beginning with mc-z: |
| Line 364: | Line 300: |
First, look at the cluster summary (or click [[FsTutorial/GroupAnalysisClusterSummary|here]]): |
First, look at the cluster summary (or click [[FsTutorial/GroupAnalysisClusterSummary|here]]): |
| Line 373: | Line 308: |
| 1. This is a list of all the clusters that were found (38 of them) 1. The CWP column is the cluster-wise probability (the number you are interested in). It is a simple p (ie, NOT -log10(p)). 1. Note that ALL clusters are found, regardless of their significance. 1. Cluster number 1 has a CWP of p=.001. 1. Cluster number 21 has a CWP of p=.077. |
1. This is a list of all the clusters that were found (38 of them) 1. The CWP column is the cluster-wise probability (the number you are interested in). It is a simple p (ie, NOT -log10(p)). 1. Note that ALL clusters are found, regardless of their significance. 1. Cluster number 1 has a CWP of p=.001. 1. Cluster number 21 has a CWP of p=.077. |
| Line 385: | Line 320: |
| }}} ---- |
}}} ---- |
| Line 390: | Line 324: |
| {{attachment:lateral_clusters.jpg}} {{attachment:medial_clusters.jpg}} Notes: 1. These are all clusters, regardless of significance. 1. When you click on a cluster, the label will tell you the cluster number (eg, cluster-021). |
{{attachment:lateral_clusters.jpg}} {{attachment:medial_clusters.jpg}} Notes: 1. These are all clusters, regardless of significance. 1. When you click on a cluster, the label will tell you the cluster number (eg, cluster-021). |
| Line 400: | Line 333: |
| 1. Find and click on cluster 1. It is lilac and has a value of -3 since this is log10(.001). The -3 is because the correlation is negative. 1. Find and click on cluster 21. Its value is -1.11 because this is log10(.077). Note that if you turn off the annotation, the cluster 21 is not visible because its significance is worse than the threshold we set (-fthresh 2, p < .01). 1. All vertices within a cluster are the same value (the p-value of the cluster). 1. You can change the cluster-wise threshold by clicking on '''Configure Overlay''' , and setting the '''Min''' value to your desired level. Alternatively, you can drag the red flag to adjust the cluster-wise threshold. As you do this clusters will appear or disappear from the surface. |
1. Find and click on cluster 1. It is lilac and has a value of -3 since this is log10(.001). The -3 is because the correlation is negative. 1. Find and click on cluster 21. Its value is -1.11 because this is log10(.077). Note that if you turn off the annotation, the cluster 21 is not visible because its significance is worse than the threshold we set (-fthresh 2, p < .01). 1. All vertices within a cluster are the same value (the p-value of the cluster). 1. You can change the cluster-wise threshold by clicking on '''Configure Overlay''' , and setting the '''Min''' value to your desired level. Alternatively, you can drag the red flag to adjust the cluster-wise threshold. As you do this clusters will appear or disappear from the surface. |
Preparations
If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs/group_analysis_tutorial cd $SUBJECTS_DIR/glm
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window (this will paste the command). Press enter to run the command.
These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into this directory. You can now skip ahead to the tutorial (below the gray line).
If You're not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
tcsh source your_freesurfer_dir/SetUpFreeSurfer.csh setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs/group_analysis_tutorial cd $SUBJECTS_DIR/glm
Notice the command to open tcsh. If you are already running the tcsh command shell, then the 'tcsh' command is not necessary. If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
Introduction
This tutorial is designed to introduce you to the "command-line" group analysis stream in FreeSurfer (as opposed to QDEC which is GUI-driven), including correction for multiple comparisons. While this tutorial shows you how to perform a surface-based thickness study, it is important to realize that most of the concepts learned here apply to any group analysis in FreeSurfer, surface or volume, thickness or fMRI.
Here are some useful Group Analysis Links:
QDEC Tutorial
FSGD Format
FSGD Examples
DODS vs DOSS
This Data Set
The data used for this tutorial is 40 subjects from Randy Buckner's lab. It consists of males and females ages 18 to 93. You can see the demographics here. You will perform an analysis looking for the effect of age on cortical thickness accounting for the effects of gender in the analysis. This is the same data set used in the QDEC Tutorial, and you will get the same result.
General Linear Model (GLM) DODS Setup
Design Matrix/FSGD File
For this example, we will model the thickness as a straight line. A line has two parameters: intercept (or offset) and a slope.
- The slope is the change of thickness with age
- The intercept/offset is interpreted as the thickness at age=0.
Parameter estimates are also called "regression coefficients".
To account for effects of gender, we will model each sex with its own line, meaning that there will be four linear parameters (also called "betas"):
- Intercept for Females
- Intercept for Males
- Slope for Females
- Slope for Males
In FreeSurfer, this type of design is called DODS (for "Different-Offset, Different-Slope").
In FreeSurfer, you can either create your own design matrices, or, if you can specify your design in terms of a FreeSurfer Group Descriptor File (FSGD), FreeSurfer will create them for you. The FSGD file is a simple text file. See this page for the format. The demographics page also has an example FSGD file for this data.
Things to do:
- Create an FSGD file for the above design. One (gender_age.fsgd) already exists so that you can continue with the exercises.
Information on how to create/view text files can be found here.
Contrasts
A contrast is a vector that embodies the hypothesis we want to test. In this case, we wish to test the change in thickness with age after removing the effects of gender. To do this, create a simple text file with the following numbers:
0 0 0.5 0.5
Notes:
- There is one value for each parameter (so 4 values)
- The intercept/offset values are 0 (nuisance)
- The slope values are 0.5 so as to average the F and M slopes
- A file called lh-Avg-thickness-age-Cor.mtx already exists
Analysis
Assemble the Data (mris_preproc)
Assembling the data simply means:
- Resampling each subject's data into a common space, and
- Concatenating all the subjects' into a single file
- Spatial smoothing (can be done between 1 and 2)
This can be done in two equivalent ways:
Previously Cached (qcached) Data
When you have run recon-all with the -qcache option, recon-all will resample data onto the average subject (fsaverage) and smooth it at various FWHM (full-width/half-max), usually 0, 5, 10, 10, 20, and 25mm. This can speed later processing. The data for this tutorial have been cached, so run:
mris_preproc --fsgd gender_age.fsgd \ --cache-in thickness.fwhm10.fsaverage \ --target fsaverage --hemi lh \ --out lh.gender_age.thickness.10.mgh
Notes:
- Only takes a few seconds because the data have been cached
- The FSGD file lists all the subjects, helps keep order.
- The independent variable is the thickness smoothed to 10mm FWHM.
- The data are for the left hemisphere
- The output is lh.gender_age.thickness.10.mgh (which already exists)
Things to do:
- Run mri_info on lh.gender_age.thickness.10.mgh to find its dimensions,ie,
mri_info lh.gender_age.thickness.10.mgh
Uncached Data
In the case that you have not cached the data, you can perform the same operations manually using the two commands below. There is no need to do this for this tutorial. OPTIONAL: THIS WILL TAKE ABOUT 10 MINUTES
mris_preproc --fsgd gender_age.fsgd \ --target fsaverage --hemi lh \ --meas thickness \ --out lh.gender_age.thickness.00.mgh
Notes:
- This resamples each subjects data to fsaverage
- Output is lh.gender_age.thickness.00.mgh, which is unsmoothed
OPTIONAL: THIS WILL TAKE ABOUT 5 MINUTES
mri_surf2surf --hemi lh \ --s fsaverage \ --sval lh.gender_age.thickness.00.mgh \ --fwhm 10 \ --cortex \ --tval lh.gender_age.thickness.10B.mgh
- This smooths by 10mm FWHM
- "--cortex" means only smooth areas in cortex (exclude medial wall). This is automatically done with qcache. You can also specify other labels.
- Output is lh.gender_age.thickness.10B.mgh.
GLM Analysis (mri_glmfit)
mri_glmfit \ --y lh.gender_age.thickness.10.mgh \ --fsgd gender_age.fsgd dods\ --C lh-Avg-thickness-age-Cor.mtx \ --surf fsaverage lh \ --cortex \ --glmdir lh.gender_age.glmdir
Notes:
- Input is lh.gender_age.thickness.10.mgh
- Same FSGD used as with mris_preproc. Maintains subject order!
- DODS is specified (it is the default)
- Only one contrast is used (lh-Avg-thickness-age-Cor.mtx) but you can specify multiple contrasts.
- "--cortex" specifies that the analysis only be done in cortex (ie, medial wall is zeroed out). Other labels can be used.
- The output directory is lh.gender_age.glmdir
- Should only take about 1min to run.
Things to do:
When this command is finished the glm directory will contain a sub-directory called lh.gender_age.glmdir. There will be a number of output files in this directory, as well as two other directories. If you did an ls in your glmdir there will be:
y.fsgd -- copy of input FSGD file (text) Xg.dat -- design matrix (text) mask.mgh -- binary mask (surface overlay) beta.mgh -- all parameter estimates (surface overlay) rstd.mgh -- residual standard deviation (surface overlay) sar1.mgh -- residual spatial AR1 (surface overlay) fwhm.dat -- average FWHM of residual (text) dof.dat -- degrees of freedom (text) lh-Avg-thickness-age-Cor -- contrast subdirectory mri_glmfit.log -- log file (text, send this with bug reports)
Study Questions:
- What was the DOF for this experiment?
- What was the FWHM?
- How many frames does beta have? Why?
There will be a subdirectory for each contrast that you specify. The name of the directory will be that of the contrast matrix file (without the .mtx extension). The lh-Avg-thickness-age-Cor directory will have the following files:
C.dat -- original contrast matrix (text) gamma.mgh -- contrast effect size (surface overlay) F.mgh -- F ratio of contrast (surface overlay) sig.mgh -- significance, -log10(pvalue), uncorrected (surface overlay)
View the uncorrected significance map with freeview:
freeview -f $SUBJECTS_DIR/fsaverage/surf/lh.inflated:annot=aparc.annot:overlay=lh.gender_age.glmdir/lh-Avg-thickness-age-Cor/sig.mgh:overlay_threshold=2,5
Choose the 3D view.
Once the aparc parcellation appears on the inflated surface, click the Show outline only checkbox to expose the uncorrected significance map.
You should see:
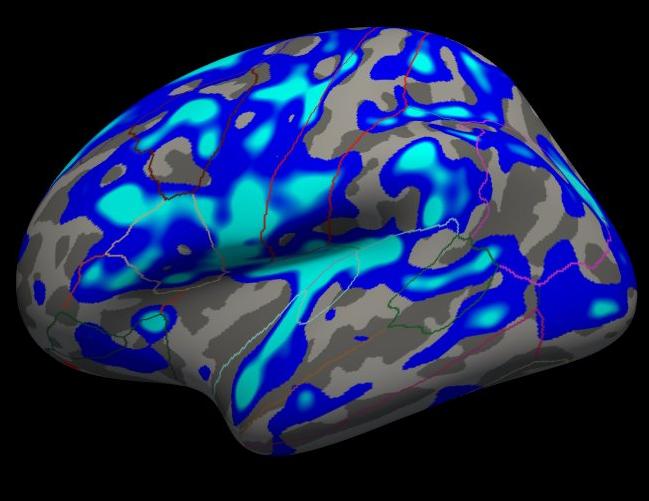
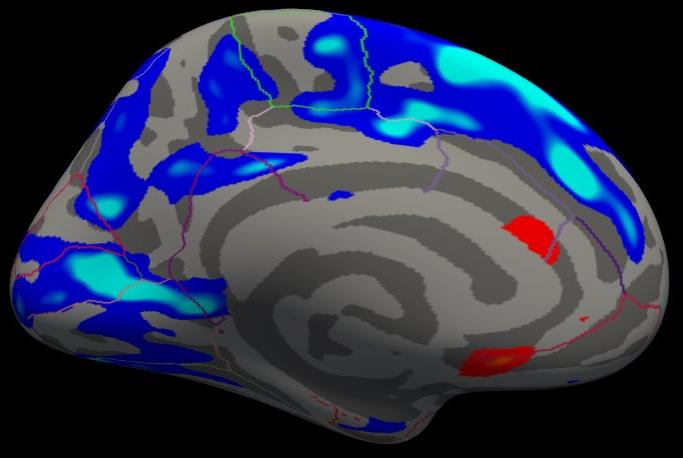
Notes:
The threshold is set to 2, meaning vertices with p<.01, uncorrected, will have color.
- Blue means a negative correlation (thickness decreases with age), red is positive correlation.
- Click on a point in the Precentral Gyrus. What is it's value? What does it mean?
Viewing the medial surface, change the overlay threshold to something very, very low (say, .01), by clicking Configure Overlay and set the Min value to 0.01. Click Apply, or check the Automatically apply changes checkbox.
You should see:
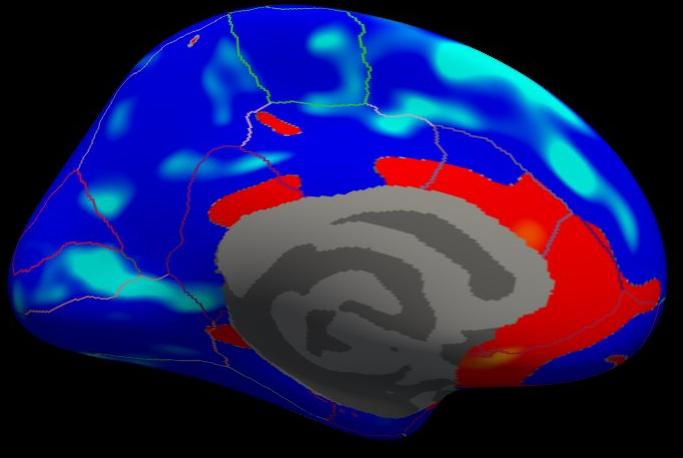
Notes:
- Almost all of cortex now has color
- The non-cortical areas are still blank (0) because they were excluded with --cortex in mri_glmfit above.
Clusterwise Correction for Multiple Comparisons
Note: The method used is based on: Smoothing and cluster thresholding for cortical surface-based group analysis of fMRI data. Hagler DJ Jr, Saygin AP, Sereno MI. NeuroImage (2006).
To perform a cluster-wise correction for multiple comparisons, we will run a simulation. The simulation is a way to get a measure of the distribution of the maximum cluster size under the null hypothesis. This is done by iterating over the following steps:
- Synthesize a z map
- Smooth z map (using residual FWHM)
- Threshold z map (level and sign)
- Find clusters in thresholded map
- Record area of maximum cluster
Repeat over desired number of iterations (usually > 5000)
In FreeSurfer, this information is stored in a simple text file called a CSD (Cluster Simulation Data).
Once we have the distribution of the maximum cluster size, we correct for multiple comparisons by:
- Going back to the original data
- Thresholding using same level and sign
- Finding clusters in thresholded map
- For each cluster, p = probability of seeing a maximum cluster that size or larger during simulation.
Run the simulation
All these steps are performed with the mri_glmfit-sim:
mri_glmfit-sim \ --glmdir lh.gender_age.glmdir \ --sim mc-z 5 4 mc-z.negative \ --sim-sign neg --cwpvalthresh 0.4\ --overwrite
Notes:
- Specify the same GLM directory (--glmdir)
- The simulation type is Z Monte Carlo (mc-z)
- Specify the sign (neg for negative, pos, or abs)
Vertex-wise threshold of 4 (p < .0001)
Only 5 iterations (usually > 5000)
- CSD files will have mc-z.negative base
"--cwpvalthresh 0.4" Keep clusters that have cluster-wise p-values < 0.4 (usually you will want something like .05). To see all clusters, set to .999
- --overwrite deletes previous CSD files
This has been run this data with 1000 iterations (CSD base of mc-z.neg). This can often take many hours.
View CSD File
The CSD files are stored in lh.gender_age.glmdir/csd. Look at one of them:
less lh.gender_age.glmdir/csd/mc-z.neg4.j001-lh-Avg-thickness-age-Cor.csd
Or click here.
To end the less command, type 'q' for quit.
Notes:
- Rows that begin with '#' are headers or comments.
- Each data row is an iteration.
- The 3rd column is the maximum cluster size for that iteration.
- The CSD file itself is not particularly important, so don't worry too much about it.
View the Corrected Results
In the contrast subdirectory, you will see several new files, all beginning with mc-z:
mc-z.neg4.sig.cluster.summary - summary of clusters (text) mc-z.neg4.sig.cluster.mgh - cluster-wise corrected map (overlay) mc-z.neg4.sig.ocn.annot - annotation of clusters
First, look at the cluster summary (or click here):
less lh.gender_age.glmdir/lh-Avg-thickness-age-Cor/mc-z.neg4.sig.cluster.summary
Notes:
- This is a list of all the clusters that were found (38 of them)
- The CWP column is the cluster-wise probability (the number you are interested in). It is a simple p (ie, NOT -log10(p)).
- Note that ALL clusters are found, regardless of their significance.
- Cluster number 1 has a CWP of p=.001.
- Cluster number 21 has a CWP of p=.077.
Load the cluster annotation in freeview
freeview -f $SUBJECTS_DIR/fsaverage/surf/lh.inflated:overlay=lh.gender_age.glmdir/lh-Avg-thickness-age-Cor/mc-z.neg4.sig.cluster.mgh:overlay_threshold=2,5:annot=lh.gender_age.glmdir/lh-Avg-thickness-age-Cor/mc-z.neg4.sig.ocn.annot
You should see:
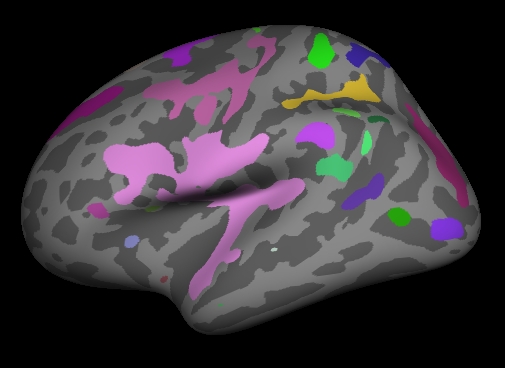
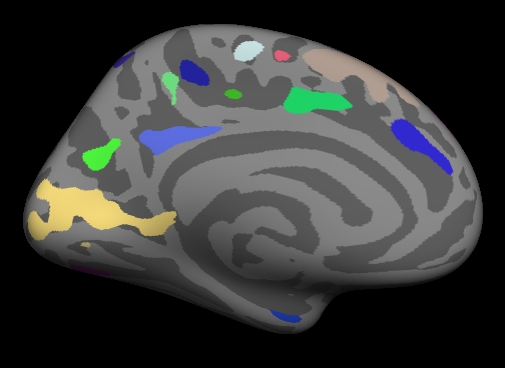
Notes:
- These are all clusters, regardless of significance.
- When you click on a cluster, the label will tell you the cluster number (eg, cluster-021).
Things to do:
- Find and click on cluster 1. It is lilac and has a value of -3 since this is log10(.001). The -3 is because the correlation is negative.
- Find and click on cluster 21. Its value is -1.11 because this is log10(.077). Note that if you turn off the annotation, the cluster 21 is not visible because its significance
is worse than the threshold we set (-fthresh 2, p < .01).
- All vertices within a cluster are the same value (the p-value of the cluster).
You can change the cluster-wise threshold by clicking on Configure Overlay , and setting the Min value to your desired level. Alternatively, you can drag the red flag to adjust the cluster-wise threshold. As you do this clusters will appear or disappear from the surface.
