Making FinalSurf edits - base
To follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names.
Edits made to the brain.finalsurfs.manedit.mgz volume of the cross sectional data sets will not transfer to the base since the base creation takes off from the norm.mgz of the cross data sets. Therefore, in order to fix the base, one must edit the brain.finalsurfs.manedit.mgz volume as follows: Below is a visualization of the problem area in the base: Coronal slice 89 at coordinates 109 166 89 in right hemi. The problem area extends through slices 85 - 91 (and zoom-in):
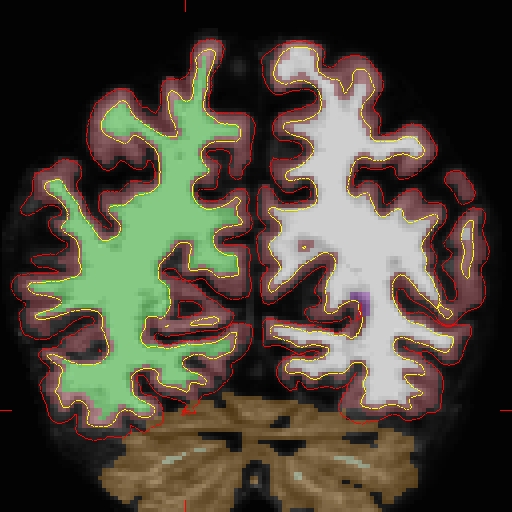
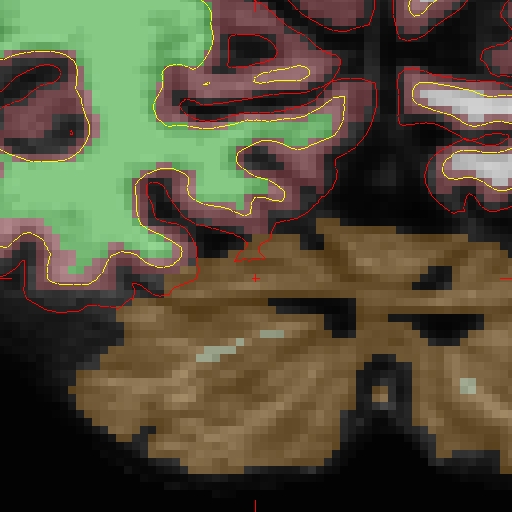
First make a copy of the brain.finalsurfs.mgz volume to one called 'brain.finalsurfs.manedit.mgz' and save it within the mri directory of the base:
cp OAS2_0002/mri/brain.finalsurfs.mgz OAS2_0002/mri/brain.finalsurfs.manedit.mgz
Next, load the brain.finalsurfs.manedit.mgz volume in Freeview (aux in tkmedit) and edit this volume by going through the affected slices and removing the voxels in the cerebellum and surrounding areas that cause the pial surface misplacement. Once you are finished editing, save the changes to the edited volume. Then, run the following to have edits take effect:
Don't run this recon-all command
It will take hours and has already been done for you.
recon-all -base OAS2_0002 -autorecon-pial
Results can be seen on OAS2_0002_fixed
freeview -v OAS2_0002_fixed/mri/brainmask.mgz \
OAS2_0002_fixed/mri/brain.finalsurfs.manedit.mgz \
OAS2_0002_fixed/mri/aseg.mgz:colormap=lut:opacity=0.25 \
-f OAS2_0002_fixed/surf/lh.pial:edgecolor=red \
OAS2_0002_fixed/surf/rh.pial:edgecolor=red \
OAS2_0002_fixed/surf/lh.white:edgecolor=blue \
OAS2_0002_fixed/surf/rh.white:edgecolor=blueand will look as follows:
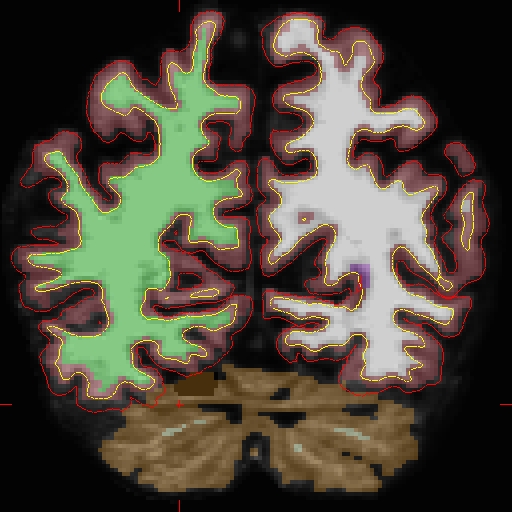
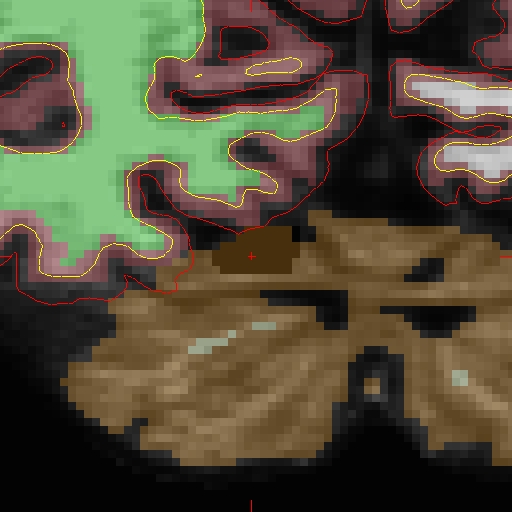
Now that the base and cross sectional time point 2 are fixed, you must re-create the longitudinal data set for time point 2. Return to the Longitudinal Tutorial to read that section.
