Back to Top Tutorial Page
Back to Multimodal Top
Other Multimodal Tutorials: A. Multimodal Registration, C. Individual fMRI Integration, D. Surface-based Group fMRI Analysis
The purpose of this tutorial is to give you experience with the integration of Diffusion Tensor Imaging (DTI) with FreeSurfer. The data were collected at MGH as part of the MIND.
DTI Basics
It is assumed that you already have knowledge of DTI and its analysis. This paragraph is supplied as a summary for completeness. DTI attempts to measure the diffusion of water in the brain. White matter tracts tend to be like little straws which constrain the direction diffusion. The DTI analysis measures the orientation and the "strength" of the orientation. Eg, CSF will have no strong orientation. This strength is often measured as the fractional anisotropy (FA). The diffusivity is often measured as the Apparent Diffusion Coefficient (ADC).
Prior to DTI analysis, it is customary to motion correct, and sometimes eddy current correct, the diffusion weighted images. This is done by selecting one of the images as a template (usually a low-b volume). This forces all the DTI-related maps to be in-line with the template. Consequently, this template should be used as the registration target.
This Data Set
This data was acquired at MGH as part of a MIND Consortium study. The raw data are in dwi.nii.gz. The FreeSurfer anatomical analysis for this subject is M87102113.v4. Seventy images were acquired (10 low-b and 60 diffusion weighted). The bvalues and gradient directions are stored in bvals.dat and bvects.dat, respectively. These data were analyzed with FreeSurfer's dt_recon program, which created the fa.nii, adc.nii, lowb.nii, and register.dat. The data are located in:
Preparations
If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/multimodal/dti
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window (this will paste the command). Press enter to run the command.
These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into this directory. You can now skip ahead to the tutorial (below the gray line).
If You're not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
tcsh source your_freesurfer_dir/SetUpFreeSurfer.csh setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/multimodal/dti
Notice the command to open tcsh. If you are already running the tcsh command shell, then the 'tcsh' command is not necessary. If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
Check the Registration
As always, check the registration with the template. In this case, the template is the low-b image:
tkregister2 --mov lowb.nii --reg register.dat --surf
The registration should be accurate. For more information on registration, see the Registration Tutorial.
FA Viewing
View the FA on the subject's anatomical with the white matter parcellation (wmparc.mgz) as the segmentation:
tkmedit M87102113.v4 orig.mgz -aux brain.mgz \ -seg wmparc.mgz \ -reg register.dat -overlay fa.nii \ -fthresh 0.2 -fmax 1
Notes:
- The FA is a values between 0 and 1, so the thresholds are set to 0.2 and 1.
- When the window first comes up, there will be a lot of activity
outside the brain. To remove this, set the 'View->Mask Functional Overlay To Aux Volume' flag.
- In the image below, you can see that white matter tends to have a higher FA.
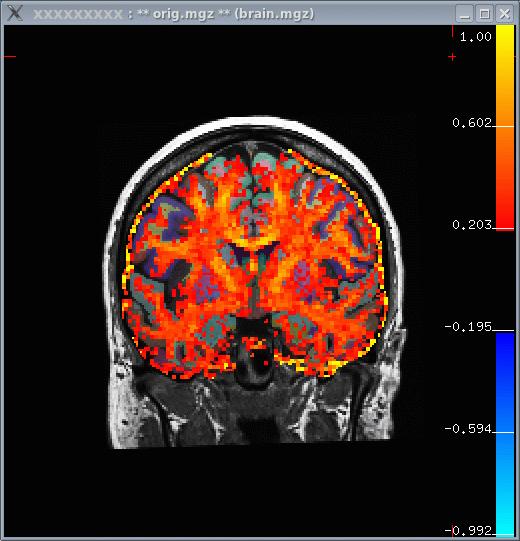
FA ROI Analysis
As with the Individual fMRI Analysis, we will first resample the FA into the subject's anatomical space:
mri_vol2vol --mov fa.nii \ --reg register.dat \ --fstarg --interp nearest \ --o fa.anat.mgh
Notes:
- Nearest neighbor is used to avoid averaging FA across voxels.
- The output fa.anat.mgh will be 256^3, 1mm3.
Now run mri_segstats getting a report for each of the following ROIs (index/name pairs are found in the LUT):
251 CC_Posterior (Posterior Corpus Callosum) 3024 wm-lh-precentral 3030 wm-lh-superiortemporal 3021 wm-lh-pericalcarine 12 Left-Putamen 4 Left-Lateral-Ventricle
The wm-lh-precentral, wm-lh-superiortemporal, wm-lh-pericalcarine are created by the gyral white matter parcellation.
Now run mri_segstats:
mri_segstats \ --seg $SUBJECTS_DIR/M87102113.v4/mri/wmparc.mgz \ --ctab $FREESURFER_HOME/FreeSurferColorLUT.txt \ --id 251 --id 3021 --id 3024 --id 3030 --id 12 --id 4 \ --i fa.anat.mgh --sum fa.stats
Click HERE to see the output. The CC has an average FA of about 0.75, gyral parcellations are about 0.4, the left putamen is 0.27, and the ventricle is 0.2. This is as expected because the CC is highly directional with no crossing fibers so we would expect the CC to have the highest FA. The gyral white matter is also directional but has fibers crossing in them, so one expects the FA to be lower than CC. The gray matter (putamen) is still lower. The ventricle has no fibers, so we expect it to have the lowest FA.
Other multimodal tutorials: A. Multimodal Registration, C. Individual fMRI Integration, D. Surface-based Group fMRI Analysis
