| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| [[FsTutorial|top]] | [[FsTutorial/TroubleshootingData|previous]] | [[FsTutorial|top]] | [[FsTutorial/TroubleshootingDataV6.0|previous]] |
Correcting Pial Surfaces
To follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names.
The pial surface is created by expanding the white matter surface so that it closely follows the gray-CSF intensity gradient as found in the brainmask.mgz volume. Once an accurate white surface is created then you can work on correcting the pial surface, if needed. The pial surface boundary and white matter surface boundary should not cross. After the pial surface has been generated, it's a good idea to visually check it for defects that may have been created during automatic topology fixing. To check the pial surface, it may be loaded into freeview and viewed along with the brainmask.mgz volume. If the surface appears not to follow the gray-CSF boundary in the volume, edits may be required.
Editing the Volume
Subject pial_edits_before has an example of the pial surface including non-cortex within the boundaries. This page will take you through fixing this and other similar problems with the pial surface including non-cortex material. First, make sure you have pial_edits_before loaded in freeview:
freeview -v pial_edits_before/mri/T1.mgz \ pial_edits_before/mri/brainmask.mgz \ -f pial_edits_before/surf/lh.white:edgecolor=yellow \ pial_edits_before/surf/lh.pial:edgecolor=red \ pial_edits_before/surf/rh.white:edgecolor=yellow \ pial_edits_before/surf/rh.pial:edgecolor=red
Use the PageUp and PageDown keys to go through the volume slice by slice (Fn+Command+Up/Down on Mac), and view the pial surface (red line) and white matter surface (yellow line). Notice the bright diagonal line in slice 157 that has caused the pial surface to expand past the actual pial boundary. This is the result of a bad segmentation incorporating a piece of the dura within the pial surface.
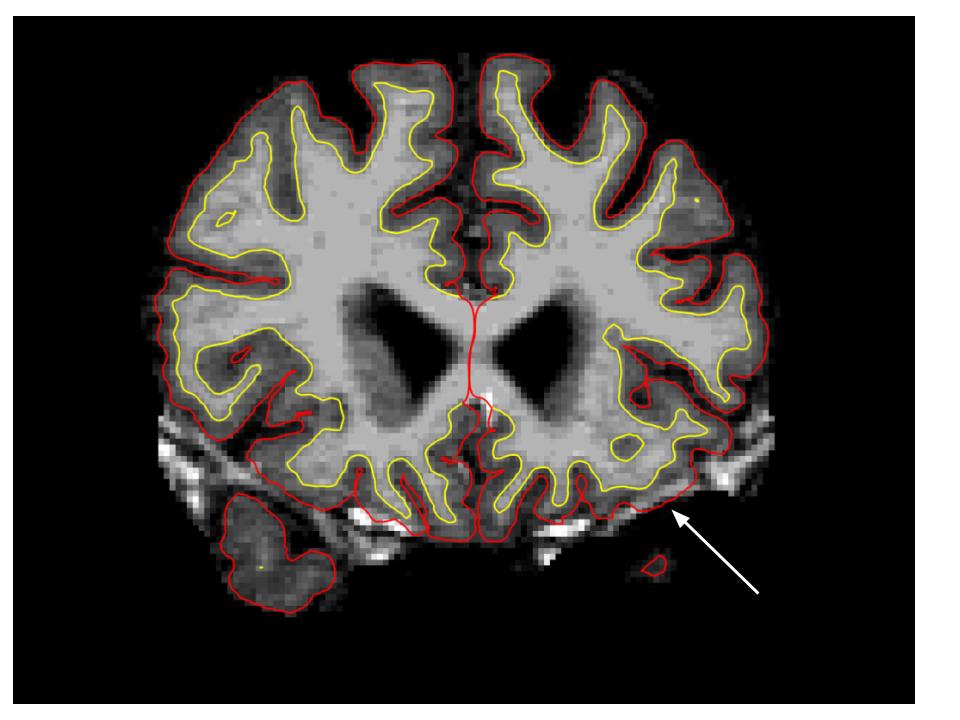
To fix this type of error you can simply edit away the offending voxels from the brainmask.mgz volume. To do this you will need to click on the Recon Edit button  which will bring up this window:
which will bring up this window:
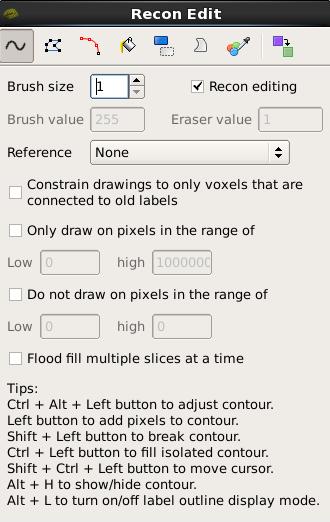
First, set the brush to a size comfortable for you. A brush size of 2 works well for this edit. The Recon editing mode is already selected for you as indicated by the checkmark in the corresponding checkbox. In Recon editing mode the brush value and eraser value are set to 255 and 1, respectively. Make sure the first option, Freehand, at the top-left, is selected and you're ready to make your edits. Find a place in the image where the dura is causing errors in the segmentation. Double check that the brainmask.mgz (not the T1.mgz) is selected and highlighted in the side panel listing all the volumes. You only want to make edits to the brainmask. Hold down the Shift key and use the left mouse button to delete the voxels. It is not necessary to completely remove the dura to get an adequate pial surface, but it is good to do so until you are more familiar with manual editing. When you are finished removing the bright diagonal line in slice 157, it should look like this:
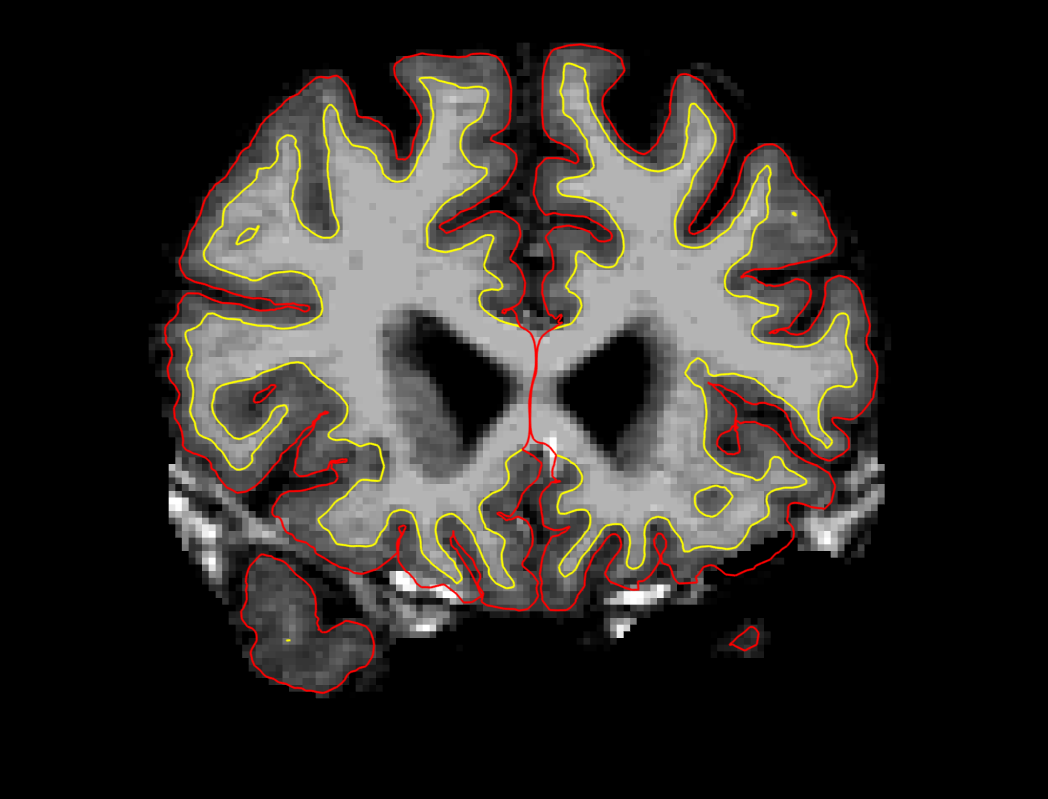
Continue on the other slices until the dura is removed. In this example, slices 156 - 160 were edited. Ctrl-Z and the Undo button  at the top in freeview allows you to go back and undo as many edits as you like. Video Tutorial:
at the top in freeview allows you to go back and undo as many edits as you like. Video Tutorial:
pialedits.mp4
At any time, you can save the changes you've made to the brainmask.mgz (if attending a course, do not press save) volume by selecting Save Volume in the freeview File menu or by clicking the Save Volume  button. Make sure the brainmask.mgz is the one highlighted while saving if you have the T1.mgz open at the same time. You can check your result by viewing the brainmask.mgz volume in the pial_edits_after directory.
button. Make sure the brainmask.mgz is the one highlighted while saving if you have the T1.mgz open at the same time. You can check your result by viewing the brainmask.mgz volume in the pial_edits_after directory.
Regenerating the Surface
When you are finished editing the voxels, you will need to regenerate the surfaces. Since the white matter hasn't been changed, you don't need to resegment the volume. You can regenerate the pial surface with:
Do not run this command if you are conducting the tutorial! This step will take a long time and there is no need to run it for the tutorial purposes.
recon-all -autorecon-pial -subjid pial_edits_before
After regenerating the pial surface, it should look like this on slice 157:
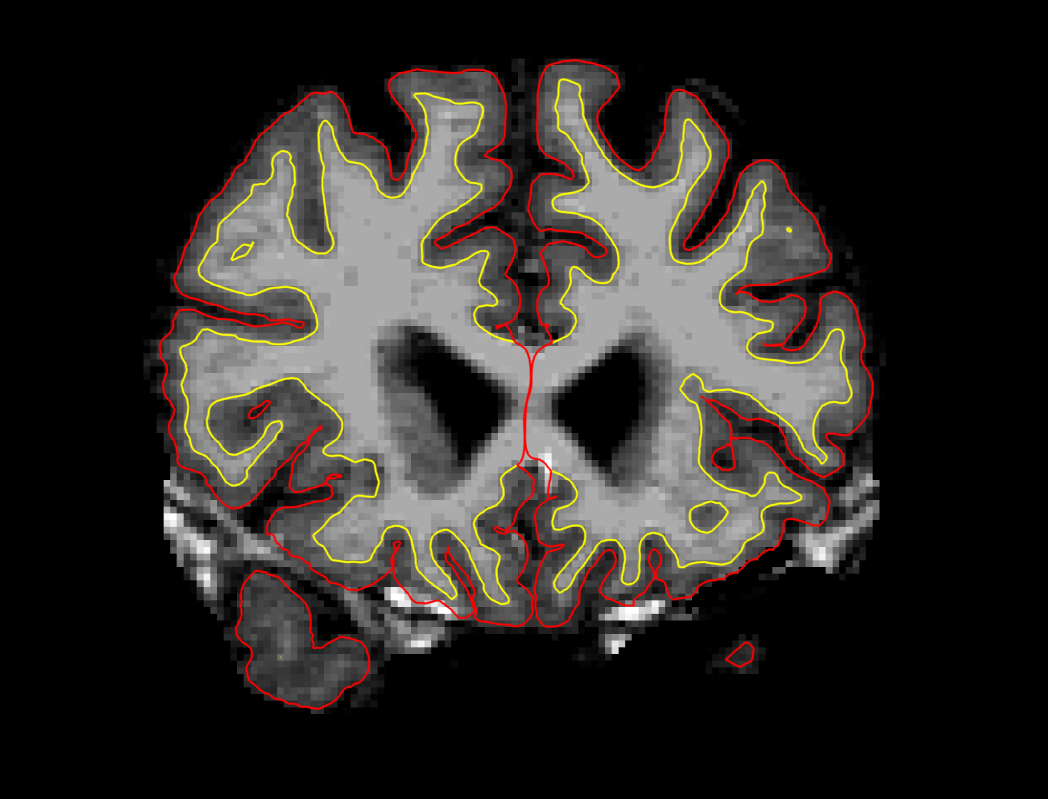
Edits to correct pial surface extension into cerebellum
If the pial surface mistakenly includes cerebellum within its boundary, you should not manually erase the cerebellum! The approach to this error is slightly different. Removing cerebellum could affect the accuracy of your aseg and future volume registrations. To correct this type of error, create a new volume called brain.finalsurfs.manedit.mgz volume and erase the cerebellum in it. To create brain.finalsurfs.manedit.mgz, you will copy it from brain.finalsurfs.mgz:
cd <subjid>/mri cp brain.finalsurfs.mgz brain.finalsurfs.manedit.mgz
Once brain.finalsurfs.manedit.mgz exists, open it in freeview and remove the cerebellum voxels that were included in the pial surface. If these are the only edits made to a subject, you would re-process the subject using the following command:
recon-all -autorecon-pial -subjid pial_edits_before
If you're interested in practicing this edit, please refer to the brain.finalsurfs section in the Longitudinal Tutorial. For all other non-cerebellum pial surface problems, brainmask.mgz should be edited.
