Contents
- Introduction
- Run FreeSurfer on Anatomical MRI
- Create a segmentation for the GTM (this is an extension of the FS recon-all analysis)
- Register your PET image with the anatomical (and motion correction if using)
- ROI Analysis, Setting Up Reference Regions for KM, and Apply Partial Volume Correction (if using)
- Kinetic Modeling (KM). KM is done with either MRTM1, MRTM2, or invasive Logan.
- Surface-based analysis
- Subcortical volume-based analysis.
- Custom Subcortical Atlases
- Custom Cortical Atlases
1. Introduction
1.1. Overview
PETSurfer provides a set of tools within FreeSurfer for end-to-end integrated MRI-PET analysis, including motion correction, PET-MRI registration, reference region kinetic modeling (MRTM1, MRTM2, Logan), partial volume correction (PVC), and group analysis in ROI, volume, and surface spaces. The kinetic modeling is made easier by integration with FreeSurfer-defined regions of interest. The PVC tools can be used in any context where PVC is needed (eg, ASL). PVC methods include the Symmetric Geometric Transfer Matrix (SGTM), two-compartment model (also known as the Meltzer Method), three-compartment model (also known as the Muller-Gartner (MG) Method), region-based voxel-wise (RBV). SGTM is used for ROI analysis where as the others are used for voxel-wise analysis. All PVC implementations also account for the tissue fraction effect (TFE).
1.2. Three-space Approach
For voxel-wise applications, PETsurfer can do standard whole-brain volumetric analysis, but the instructions below use the "Three-Space Approach". In this approach, the cortical and subcortical GM are analyzed separately (cortex is analyzed in the left and right hemispheres separately making the number of spaces equal to 3). It is done in this way to be able to exploit surface-based operations for cortical regions. This includes surface-based (2D) smoothing, which allows spatial smoothing without exacerbating partial volume effects as when 3D volumetric smoothing is used. This improves kinetic modeling significantly but helps with uptake studies. When performing PVC with MG, it is necessary to use a surface-based approach as MG with 3D smoothing is invalid. The three spaces can be recombined into a more traditional volume space for visualization. The three-space approach adds complexity, but the improved results are worth it.
1.3. How to cite
If you use PETsurfer, please cite these papers:
Greve, D. N., Svarer, C., Fisher, P. M., Feng, L., Hansen, A. E., Baare, W., ... & Knudsen, G. M. (2014). Cortical surface-based analysis reduces bias and variance in kinetic modeling of brain PET data. Neuroimage, 92, 225-236.
Greve, D. N., Salat, D. H., Bowen, S. L., Izquierdo-Garcia, D., Schultz, A. P., Catana, C., ... & Johnson, K. A. (2016). Different partial volume correction methods lead to different conclusions: An 18 F-FDG-PET study of aging. NeuroImage, 132, 334-343.
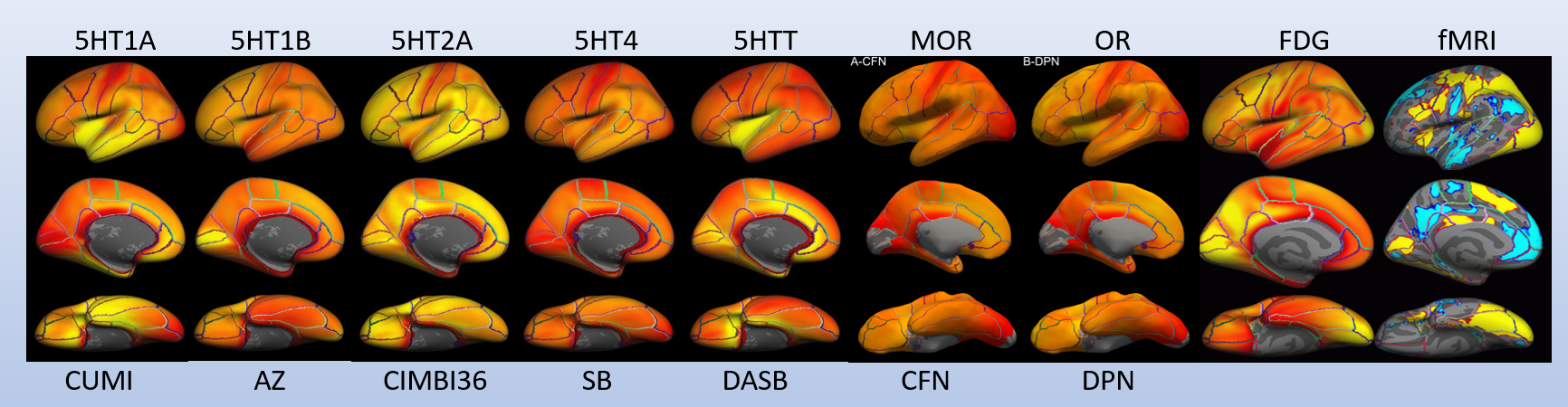
1.4. Example PET-based atlases (group averages) created by PETsurfer (plus an fMRI map)
2. Run FreeSurfer on Anatomical MRI
In all cases, you will need a T1-weighted MRI of your subject of sufficient quality to run in FreeSurfer. This does require that you know enough about FreeSurfer to run the anatomical analysis. The FreeSurfer analysis (eg, recon-all -s subject -i T1.mri.nii.gz -all) must be done first; you should QA the FS analysis before proceeding with PETsurfer. If possible, perform MRI gradient distortion correction (grad_unwarp) of the MRI before running recon-all; on a Siemens system, you can get the spherical harmonics table from C:\Medcom\MriSiteData\GradientCoil\coeff.grad (this is Siemens proprietary, so don't distribute it!).If you want to use the CVS non-linear volumetric registration, then run mri_cvs_register --mov subject --template cvs_avg35_inMNI152 after you are completely finished with the anatomical analysis; this may take an additional 12 hours or so. After that, follow the steps below.
3. Create a segmentation for the GTM (this is an extension of the FS recon-all analysis)
gtmseg --s subject
where "subject" is the name of the FreeSurfer subject when you ran recon-all. This creates a high-resolution segmentation (gtmseg.mgz) in the FS folder used to run the PVC methods. This should take about an hour or two. gtmseg.mgz will use aseg.mgz for subcortical structures, ?h.aparc.annot for cortical structures, and will estimate some extra-cerebral structures. There are ways to customize this segmentation to use different ROI definitions (eg, aparc.a2009s instead of aparc). This will create a file called $SUBJECTS_DIR/subject/mri/gtmseg.mgz
4. Register your PET image with the anatomical (and motion correction if using)
First, create a template image. If your PET data only has one frame (eg, an uptake image), then that will be your template. If your PET data has multiple frames (ie, dynamic), then you will need to create the template from the dynamic data. This can be done by extracting a single frame (mri_convert pet.nii.gz --frame frameno template.nii.gz) or averaging all the time frames together (eg, mri_concat pet.nii.gz --mean --o template.nii.gz). The way the run the rigid (6 DOF) registration is done depends a little on the nature of the PET uptake. If there is little uptake outside of the brain (eg, FDG and many receptor ligands), then this command line can be used:
mri_coreg --s subject --mov template.nii.gz --reg template.reg.lta
If there is a lot of uptake outside of the brain (eg, tau, tspo), then use this command:
mri_coreg --s subject --mov template.nii.gz --reg template.reg.lta --ref nu.mgz --no-ref-mask
For parallel operation, add --threads N to the mri_coreg command where N is the number of CPUs you want to use.
You should check the registration with:
tkregisterfv --mov template.nii.gz --reg template.reg.lta --surfs
If you are not using PVC, you can use the template.reg.lta to sample the PET volume onto the surface using mri_vol2surf, then apply standard surface-based analysis.
If you need to do motion correction, you can run
mc-afni2 --template template.nii.gz --i pet.nii.gz --o pet.mc.nii.gz --mcdat motion-parameters.dat
Then use pet.mc.nii.gz instead of pet.nii.gz below
5. ROI Analysis, Setting Up Reference Regions for KM, and Apply Partial Volume Correction (if using)
mri_gtmpvc --i pet.nii.gz --reg template.reg.lta --psf FWHM --seg gtmseg.mgz --default-seg-merge --auto-mask 1 .01 --mgx .01 --o gtmpvc.output
--psf FWHM is the full-width/half-max of the the point-spread function (PSF) of the scanner as measured in image space (also known as the burring function). The blurring function depends on the scanner and reconstruction method; here it is modeled as an isotropic Gaussian filter of the given FWHM. Eg, an HR+ is typically about 6mm. This parameter has nothing to do with applying smoothing. Set this to 0 to turn off PVC. If you don't know what to set this to, ask your PET physicist.
--seg gtmseg.mgz is the segmentation created with gtmseg
--default-seg-merge will merge several segmentations, eg, all the ventricular CSF segments will be merged into one ROI
--auto-mask 1 .01 will create a mask to exclude voxels from the analysis (saves time). Output volumes will be reduced to a bounding box around this mask (saves space)
--mgx .01 Run Muller-Gartner analysis. 01 is the GM threshold. Only necessary if you want to do a voxel-wise analysis.
--o gtmpvc.output This is the output folder.
If you do not want to perform PVC, you can just run
mri_gtmpvc --i pet.nii.gz --reg template.reg.lta --seg gtmseg.mgz --default-seg-merge --auto-mask 1.01 --no-tfe --o gtmpvc.output
The --no-tfe will turn off correction for the tissue fraction effect (TFE), which is a type of PVC. If you want to save the output volume rescaled, then add --save-input
There will be many files in the output folder some of which are described here:
gtm.stats.dat is an easy to read text file where each row is an ROI, something like:
9 17 Left-Hippocampus subcort_gm 473 174.083 1.406 0.1216
9 = ninth row
17 = index for ROI
Left-Hippocampus = name of ROI
subcort_gm = tissue class
473 = number of PET voxels in the ROI
174 = variance reduction factor for ROI (based on GLM/SGTM)
1.406 = PVC uptake of ROI relative to Pons
0.1216 = resdiual varaince across voxels in the ROI
By default, this will scale by the intensity in pons. If you do not want scaling (eg, when doing a dynamic analysis), add --no-rescale
gtm.nii.gz is a nifti file with each "voxel" being an ROI. The value is the PVC uptake of ROI relative to Pons. For single time point data, this will be totally redundant with gtm.stats.dat. Use of the NIFTI format makes it easy to concatenate (mri_concat) these files together for group analysis (mri_glmfit).
nopvc.nii.gz - same interpretation as gtm.nii.gz except that the values have not been PVCed in any way. If you are not performing PVC, then this will be the same aseg gtm.nii.gz
mgx.{ctxgm,subctxgm,gm} - these are the voxel-wise values corrected using the "extended" Muller-Gartner method. ctxgm is cortical GM, subctxgm is the subcortical GM, and gm is all GM. These volumes will be of the same resolution as the input PET but the field of view will be reduced to that of a bounding box around the mask.
aux - this subfolder has auxiliary data that are often not of immediate use to the user. The exception is bbpet2anat.lta. This is a registration file that can be used to map the output PET volume (in the mask bounding box) to the anatomical space. This file should be used when mapping the volume to the surface, etc.
If you are doing dynamic (kinetic) analysis, then add the following arguments when running mri_gtmpvc:
--km-ref 8 47 --km-hb 11 12 13 50 51 52 --no-rescale
--km-ref 8 47 specifies the ROIs to use as the reference region for the MRTM models. 8 and 47 are the cerebellar hemisphers as found in $SUBJECTS_DIR/subject/mri/gtmseg.ctab (see also $FREESURFER_HOME/FreeSurferColorLUT.txt). This creates the file km.ref.tac.dat with the reference value for each time point.
--km-hb 11 12 13 50 51 52 specifies the ROIs to use as the high-binding region if using MRTM2. This creates km.hb.tac.nii.gz with the value for the high-binding region for each time point.
6. Kinetic Modeling (KM). KM is done with either MRTM1, MRTM2, or invasive Logan.
To run MRTM1 (which must be run prior to MRTM2):
mri_glmfit --y km.hb.tac.nii.gz --mrtm1 km.ref.tac.dat time.dat --o mrtm1 --no-est-fwhm --nii.gz
where time.dat is a simple ASCII text file with the acquisition time (in seconds) of each time point in the time activity curve (TAC). This will create a folder called mrtm1 in which will be a file called k2prime.dat. The value of k2prime will be used in the MRTM2 analysis below
For the MRTM2 analysis
set k2p = `cat mrtm1/k2prime.dat` mri_glmfit --y gtm.nii.gz --mrtm2 km.ref.tac.dat time.dat $k2p --o mrtm2 --no-est-fwhm --nii.gz
This will create a folder called mrtm2 in which the bp.nii.gz will be the non-displaceable binding potential (BPND) for each ROI.
For Logan,
mri_glmfit --y gtm.nii.gz --logan blood.tac.dat time.dat tstar --o logan --no-est-fwhm --nii.gz
where tstar is the time to equilibration in seconds. For Ichise 2002 MA1-Logan,
mri_glmfit --y gtm.nii.gz --logan-ma1 blood.tac.dat time.dat tstar --o logan --no-est-fwhm --nii.gz
where tstar is the time to equilibration in seconds.
7. Surface-based analysis
Details of surface-based analysis are available in other locations in the wiki, but here is basically what you do.
If you are not using PVC, then
mri_vol2surf --mov pet.nii.gz --reg template.reg.lta --hemi lh --projfrac 0.5 --o lh.pet.fsaverage.sm00.nii.gz --cortex --trgsubject fsaverage
This command samples it onto the surface of fsaverage via the individual subject's surface. --cortex says to mask out non-cortical regions in the medial wall. --projfrac 0.5 says to sample halfway between the white and pial surfaces (ie, the middle of cortex). If you want to average over the cortical ribbon, you can use --projfrac-avg .2 .8 .1, which says to start 20% into the ribbon, sample every 10%, and stop at 80% of the thickness. pet.nii.gz is your input pet volume. The output is the lh.pet.fsaverage.sm00.nii.gz, which is a surface overlay.
When using PVC, the mgx.ctxgm.nii.gz is used, but the commands will apply to any of the volumes in the GTM output folder. Sample the mgx volume onto the left hemisphere surface
mri_vol2surf --mov mgx.ctxgm.nii.gz --reg aux/bbpet2anat.lta --hemi lh --projfrac 0.5 --o lh.mgx.ctxgm.fsaverage.sm00.nii.gz --cortex --trgsubject fsaverage
This is the same command as for no PVC, just changing the input to be the MGX volume and changing the name out of the output surface overlay.
If you are not doing kinetic modeling, concatenate all your subjects together into a stack file
mri_concat subj1/lh.mgx.ctxgm.fsaverage.sm00.nii.gz subj2/lh.mgx.ctxgm.fsaverage.sm00.nii.gz ... --o all.lh.mgx.ctxgm.fsaverage.sm00.nii.gz --prune
For no PVC, just use the lh.pet.fsaverage.sm00.nii.gz files instead.
Smooth on the surface
mris_fwhm --smooth-only --i all.lh.mgx.ctxgm.fsaverage.sm00.nii.gz --fwhm 5 --o all.lh.mgx.ctxgm.fsaverage.sm05.nii.gz --cortex --s fsaverage --hemi lh
For no PVC, just use the all.lh.pet.fsaverage.sm00.nii.gz files instead and call the output all.lh.pet.fsaverage.sm05.nii.gz
Run group analysis GLM
mri_glmfit --y all.lh.mgx.ctxgm.fsaverage.sm05.nii.gz --surface fsaverage lh --fsgd your.fsgd ...
For no PVC, just use the all.lh.pet.fsaverage.sm05.nii.gz
Note: when running correction for multiple comparisons (mri_glmfit-sim), it is highly recommended that permutation be used.
For the kinetic modeling MRTM2 surface-based analysis, lh.mgx.ctxgm.fsaverage.sm00.nii.gz (or No PVC equivalent) is the surface-based TAC for the individual sampled onto fsaverage. Smooth (mris_fwhm) this file rather that concatenating across subject. Then use the smoothed file as input to the MRTM2 analysis:
set k2p = `cat mrtm1/k2prime.dat` mri_glmfit --y lh.mgx.ctxgm.fsaverage.sm05.nii.gz --mrtm2 km.ref.tac.dat time.dat $k2p --o mrtm2.lh.sm05 --no-est-fwhm --nii.gz --surface fsaverage lh
This will create mrtm2.lh.sm05/bp.nii.gz.
These can be concatenated across subject, eg,
mri_concat subj1/mrtm2.lh.sm05/bp.nii.gz subj2/mrtm2.lh.sm05/bp.nii.gz ... --o all.mrtm2.lh.sm05.bp.nii.gz --prune
Do not perform another round of smoothing on this output. If you want more smoothing, redo the smoothing of the TACs.
Then run group analysis with mri_glmfit with all.mrtm2.lh.sm05.bp.nii.gz as input.
mri_glmfit --y all.mrtm2.lh.sm05.bp.nii.gz --surface fsaverage lh --fsgd your.fsgd
8. Subcortical volume-based analysis.
8.1. Resample into the volumetric atlas space (MNI305 or MNI152)
Note: these commands will mask to just subcortical GM (cortical GM is analyzed on the surface). They are shown with the MGX volume, but you can use the input too.
Option 1: Sample into the 2mm space of the MNI305 with an AFFINE transform
mri_vol2vol --mov subctxgm.nii.gz --reg aux/bbpet2anat.lta --tal --talres 2 --o subctxgm.mni305.2mm.sm00.nii.gz
When not using PVC, use --mov pet.nii.gz --reg template.reg.lta instead.
Option 2: Sample into the 2mm space of the MNI152 with the non-linear CVS transform
mri_vol2vol --gcam subctxgm.nii.gz aux/bbpet2anat.lta $SUBJECTS_DIR/$subject/cvs/final_CVSmorph_tocvs_avg35_inMNI152.m3z $FREESURFER_HOME/subjects/cvs_avg35_inMNI152/mri.2mm/register.lta 0 1 subctxgm.cvs.2mm.sm00.nii.gz
When not using PVC:
mri_vol2vol --gcam pet.nii.gz template.reg.lta $SUBJECTS_DIR/$subject/cvs/final_CVSmorph_tocvs_avg35_inMNI152.m3z $FREESURFER_HOME/subjects/cvs_avg35_inMNI152/mri.2mm/register.lta 0 1 subctxgm.cvs.2mm.sm00.nii.gz
8.2. Smooth in 3D within SubCortical GM
This command shows 5mm, but it could be another value. Also, the commands below show mni305, but it could be cvs instead. Substitute the No PVC volume as needed.
mri_fwhm --smooth-only --i subctxgm.mni305.2mm.sm00.nii.gz --fwhm 5 \ --o subctxgm.mni305.2mm.sm05.nii.gz \ --mask $FREESURFER_HOME/subjects/fsaverage/mri.2mm/subcort.mask.mgz
If not doing kinetic modeling, then the subctxgm.mni305.2mm.sm05.nii.gz can be mri_concat together for group analysis.
If doing KM, then
set k2p = `cat mrtm1/k2prime.dat` mri_glmfit --y subctxgm.mni305.2mm.sm05.nii.gz --mrtm2 km.ref.tac.dat time.dat $k2p --o mrtm2.subctxgm.mni305.2mm.sm05 --no-est-fwhm --nii.gz
This will create mrtm2.mni305.2mm.sm05/bp.nii.gz. These can be concatenated across subject, eg,
mri_concat subj1/mrtm2.mni305.2mm.sm05/bp.nii.gz subj2/mrtm2.mni305.2mm.sm05/bp.nii.gz ... --o all.mrtm2.mni305.2mm.sm05.bp.nii.gz --prune
Do not perform another round of smoothing on this output. If you want more smoothing, redo the smoothing of the TACs.
Then run group analysis with mri_glmfit with all.mrtm2.mni305.2mm.sm05.bp.nii.gz as input.
mri_glmfit --y all.mrtm2.mni305.2mm.sm05.bp.nii.gz --fsgd your.fsgd ...
Note: when running correction for multiple comparisons (mri_glmfit-sim), it is highly recommended that permutation be used.
9. Custom Subcortical Atlases
By default, gtmseg will use the subcortical structure in aseg.mgz. This can be customized if you have a specific segmentation (eg, internal globus pallidum (GPi)). It is assumed that you will have an mgz volume with the GPi segmentation labeled in it. You will need to create a color table for your segmentation, eg,
# TissueTypeSchema #ctTType 0 unknown 0 0 0 0 #ctTType 1 cortex 205 62 78 0 #ctTType 2 subcort_gm 230 148 34 0 #ctTType 3 wm 0 255 0 0 #ctTType 4 csf 120 18 134 0 #ctTType 5 head 150 150 200 0 268 Left-GPi 219 100 176 0 2 269 Right-GPi 219 100 176 0 2
This color table has a tissue-type map in it. Left and Right GPi are given indices 268 and 269. The three numbers to the right (219 100 176) are the red-green-blue weights that dictate what color the segmentation will be in freeview. The next number is always 0. The final number (2) indicates which tissue type the structure is in as indicated in the TissueTypeSchema; in this case, GPi is a subcortical gray matter structure so it gets 2 (subcort_gm). The indices should be unique with respect to the indices in the aseg (eg, don't use 17 because is hippocampus). Next, run gtmseg to create the apas+head.mgz file. Then run
cd $SUBJECTS_DIR/subject/mri mergeseg --src apas+head.mgz --merge yourseg.mgz --o apas+head+yourseg.mgz gtmseg --s subject --head apas+head+myseg.mgz --ctab myseg.colortable.txt --o gtmseg+myseg.mgz
This will create gtmseg+myseg.mgz and gtmseg+myseg.ctab; note that this can be combined with custom cortical atlases as below. The output can be loaded into freeview with
freeview orig.mgz gtmseg+myseg.mgz:colormap=lut:lut=gtmseg+myseg.ctab
Inspect this for at least one subject to make sure that your segmentations are showing up in the right place with the right color and the right name. Look in gtmseg+myseg.ctab to confirm that the tissue type is correct. When running mri_gtmpvc, just specify --seg gtmseg+myseg.mgz
10. Custom Cortical Atlases
By default, gtmseg will use the ?h.aparc.annot cortical atlas. If you have your own atlas or you want to use another atlas inside of FreeSurfer (eg, ?h.aparc.a2009s.annot), you can do so by simply specifying
gtmseg --s subject --ctx-annot annotname lhbase rhbase --o gtmseg+myannot.mgz
where annot name is the name of the annotation (eg, aparc.a2009s; no need to include the .annot). The lhbase and rhbase will dictate the segmentation indices in the output volume. The annot will have a color table embedded in it in which each label will be given an index. For left hemi labels, this index will be added to lhbase to generate the index for the volume. Eg, --ctx-annot aparc 1000 2000, then lh pericalcarine will be 21+1000=1021; for rh it will be 21+2000. This will apear in the output gtmseg colortable. There are two constraints on how you set the base values. First, they must be 1000 or greater so that downstream software knows that this is cortical gray matter. Second, the difference between the bases must be greater than the number of cortical labels, otherwise the lh and rh indicies will overlap. The new labels should appear in the gtmseg+myannot.ctab as cortex tissue type. Load the output into freeview (see above) to confirm that the segmentation is correct. When you run mri_gtmpvc, specify --seg gtmseg+myannot.mgz.
