| Deletions are marked like this. | Additions are marked like this. |
| Line 3: | Line 3: |
| [[TableOfContents]] | <<TableOfContents>> |
| Line 16: | Line 16: |
| attachment:gw_ca_label.gif | {{attachment:gw_ca_label.gif}} |
| Line 31: | Line 31: |
| attachment:gw_make_label_atlas.gif | {{attachment:gw_make_label_atlas.gif}} |
The Automatic Surface Labeling Process ; GC Atlases
Index
Contents
Overview
This document provides an overview of two related topics:
How the FreeSurfer pipeline automatically labels a subject's regions (producing an annotation file) by comparing the subject's curvature data to an "atlas" ("gcs") file. The surface-region-labeling is sometimes called "parcellation".
The process of creating one's own atlas file, perhaps to define regions different from the ones mapped in the atlases supplied with FreeSurfer.
The Labeling Process
First, the labeling process depends on the subject having already been through the spherical registration process (ie: having a ?h.sphere.reg file), because the atlas data is in the same space as the registration template. This is the case for the recon-all FreeSurfer pipeline script. (The separate topic of spherical registration is described here: SurfaceRegAndTemplates.)
The labeling process itself is diagrammed below.
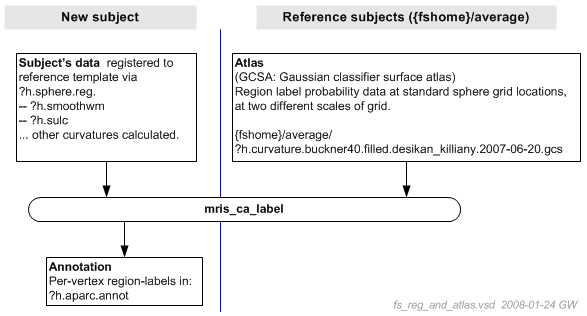
At first glance, mris_ca_label might appear to simply cross-reference a subject's vertex positions to atlas space and copy the labels of the nearest atlas points into an annotation file for the subject. However, the process is more sophisticated.
The atlas data captures not just the most likely label for a particular atlas location. Instead, the atlas data includes the probability of a particular label given values for other measures, such as convexity and curvature of the smoothwm surface. This probability data is captured in a form refered to as "gaussian classifiers", collections of parameters at each point in a regular spherical grid. (Actually two sets of gaussian classifiers at two different grid resolutions.)
Hence, in addition to the subject's ?h.sphere.reg file, mris_ca_label inputs the subject's convexity data (?h.sulc) and ?h.smoothwm surface (from which it calculates more curvature measures). This data is compared to the atlas GCs to determine the most likely label for each subject surface vertex.
Creating One's Own Atlas
To prepare to automatically label regions differently from those provided by the supplied atlases, it might be desirable to create one's own atlas. (In a few circumstances, it might also be desirable to develop a new surface registration template; see: SurfaceRegAndTemplates.)
To create an atlas, the labor-intensive first step is to manually create labels for the regions you want, on a group of reference subjects. These can be drawn using tksurfer, resulting in your own custom annotation file for each of the reference subjects. In some cases, it may save time to start with existing annotation files and edit them, rather than starting from scratch. (For more on labels and annotation files: LabelsClutsAnnotationFiles).
With custom annotation files completed, program mris_ca_train is used, as diagrammed below.
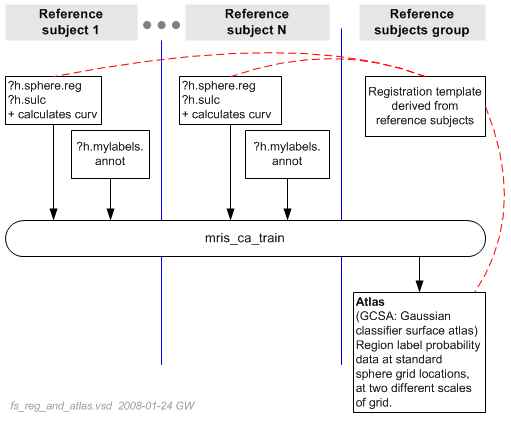
mris_ca_train reads data from each subject, including the manual annotation and combines it across subjects to create the atlas file. The dotted red lines are a reminder that implicitly the registration template, though not directly used in this process, is what underlies the registration of atlas to the reference subjects.
(The same reference subjects might be used to create the registration template and the manual annotations for the region atlas. Alternatively, new reference subjects could be used for the atlas, registered to a template previously created with an earlier group of reference subjects. The latter describes the case where you register your subjects using FreeSurfer's provided template, but then devise your own atlas by manually annotating a group of your own subjects.)
References
Automatic parcellation is the major theme of:
- Automatically Parcellating the Human Cerebral Cortex, Fischl, B., A. van der Kouwe, C. Destrieux, E. Halgren, F. Segonne, D. Salat, E. Busa, L. Seidman, J. Goldstein, D. Kennedy, V. Caviness, N. Makris, B. Rosen, and A.M. Dale, (2004). Cerebral Cortex, 14:11-22.
An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest, Desikan, R.S., F. Segonne, B. Fischl, B.T. Quinn, B.C. Dickerson, D. Blacker, R.L. Buckner, A.M. Dale, R.P. Maguire, B.T. Hyman, M.S. Albert, and R.J. Killiany, (2006). NeuroImage 31(3):968-80.
