Contents
- Introduction
- Installation
- Basic Single Input Usage
- Multiple Inputs
- Table Output (--write_volumes)
- Quality Assurance (--write_qa_stats)
- Including Estimated Intracranial Volume (eTIV, --etiv)
- Group GLM Analysis
- Running in FreeSurfer Mode
- Input with voxel sizes that are not 1mm Isotropic (--conform)
- Other options
- Source Code
- U-Net Training and Architecture
Introduction
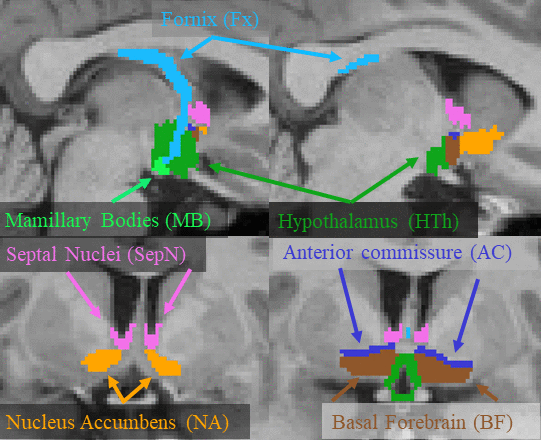
This page describes a deep learning tool, mri_sclimbic_seg, to automatically segment several subcorical limbic structures from T1-weighted images ("SC" = subcortical). The structures are shown in the figure below (note: anterior commissure is only present for reference reasons). Note there is another tool to segment the hypothalamic subunits.
Those using this toolbox should cite: A Deep Learning Toolbox for Automatic Segmentation of Subcortical Limbic Structures from MRI Images. Greve, DN, Billot, B, Cordero, D, Hoopes, A, Hoffmann, M, Dalca, A, Fischl, B, Iglesias, JE, Augustinack, JC. Submitted to Neuroimage 2021.
Installation
Currently, you can obtain ScLimbic in two ways. One is to download the development version of FreeSurfer or you can download a "stripped-down" version (linux and mac) that has just what you need to run sclimbic, view results, and run group analysis. You do not need to run or understand FreeSurfer to run the basic ScLimbic, though ScLimbic can interface with FreeSurfer analysis. For the stripped-down version, you need to set it up with
cd /path-to-install tar xvfz sclimbic-linux-20210725.tar.gz export FREESURFER_HOME=/path-to-install/sclimbic source $FREESURFER_HOME/setup.sh
If you are in the Martinos Center, you can gain access by sourcing the development environment: source /usr/local/freesurfer/nmr-dev-env
Basic Single Input Usage
In the basic usage, one passes the T1w volume (NIFTI or mgz format) one wants to segment and specifies the output:
mri_sclimbic_seg --i bert.mgz --o bert.sclimbic.mgz
This should take less than 1min to run. If the resolution is not 1mm3, then you must add the --conform. The segmentation can be viewed with freeview or tkmeditfv:
freeview bert.mgz bert.sclimbic.mgz
and should look like the figure above.
Note: if you have the stripped-down version, you can find bert in $FREESURFER_HOME
Note: if you are using NIFTI images, then you need to run freeview bert.mgz bert.sclimbic.nii:colormap=lut:lut=$FREESURFER_HOME/models/sclimbic.ctab
Multiple Inputs
If one has multiple volumes, one can create a folder with the T1 volumes in (NIFTI or mgz format) one wants to segment, then run:
mri_sclimbic_seg --i inputfolder --o outputfolder --write_volumes --write_qa_stats
The tool will find all the input images, segment them, and write out the segmentation volumes into the output folder. If the images are named subject1.mgz subject2.mgz, etc, the segmentations will be named subject1.sclimbic.mgz, subject2.sclimbic.mgz, etc. The other options (--write_volumes --write_qa_stats) are explained below.
Table Output (--write_volumes)
Oftentimes, one wants to have the volume (in mm3) of the structures, eg, for use in a group analysis. To have ScLimbic produce a table for you, add --write_volumes to the command line:
mri_sclimbic_seg --i inputfolder --o outputfolder --write_volumes
The above command will also create a CSV file called sclimbic_volumes_all.csv where each row is a case, each column is a label, and each entry is the volume of that structure in mm3.
Quality Assurance (--write_qa_stats)
It is important to check the quality of the segmentation as U-Nets can perform poorly when they see inputs that are very different than that they were trained on. The best way to check the data is to simply visualize it using the freeview command above. However, this can be tedious when there are many subjects. Accordingly, ScLimbic can produce two QA files when run with --write_qa_stats:
mri_sclimbic_seg --i inputfolder --o outputfolder --write_volumes --write_qa_stats
This will produce two other CSV files, sclimbic_zqa_scores_all.csv and sclimbic_confidence_all.csv. The ZQA is a z-score computed based on the manually labeled subjects that went into the training. The "confidence" is the mean posterior probability within the label. If the z-score is very high or the confidence is very low, then the case should definitely be visually examined. The z-score should only be used for quality control. It should not be interpreted as a deviation from a normal population because the subjects used for manual labeling were not chosen with this in mind. Your results may have systematically larger or smaller volumes than the average in the training set; this is ok.
Including Estimated Intracranial Volume (eTIV, --etiv)
If one is going to perform a group analysis on the structure volume (eg, comparing subjects with Alzheimer's against healthy controls), then one should corrected for head size since the brain size scales with head size. If you do not have your own measure, ScLimbic will compute one for you based on the method used in FreeSurfer based on Buckner, et al, 2004, NeuroImage 23:724-738. The simple way to do this is to just add --etiv to the mri_sclimbic_seg command line:
mri_sclimbic_seg --i inputfolder --o outputfolder --write_volumes --write_qa_stats --etiv
Note: this will increase the analysis time for each subject to about 5min -- but, hey, it's still pretty fast. When this is done, you will see an eTIV column in the CSV file. This will be automatically done when using ScLimbic in FreeSurfer mode (see below).
Group GLM Analysis
When performing a group analysis (eg, comparing the mean volume of one population against another and/or regress against a variable), one can simply load the CSV into your statistical analysis program (eg, R, SPSS, matlab). Alternatively, one can use the native tool in FreeSurfer called mri_glmfit on the output table, eg,
mri_glmfit --table sclimbic_volumes_all.csv --fsgd group.fsgd --scale-by-etiv --o glm.results
where group.fsgd is a FreeSurfer Group Descriptor File where the groups, regressors, and contrasts are defined. See Examples. Adding --scale-by-etiv requires that the CSV file has an eTIV column and causes mri_glmfit to scale the volumes by 100,000/eTIV (ie, the result is one-thousandth of a percent of eTIV). The main outputs will be: glm.results/sig.table.dat and glm.results/gamma.table.dat. The sig file will be the significance values stored as -log10(p) and signed by the sign of the contrast (eg, if p=0.01, then sig == -log10(.01) = 2). The gamma file will be the contrast value.
Running in FreeSurfer Mode
Instead of using the --i flag to provide input, ScLimbic can take input from subjects already run through FreeSurfer's recon-all. To do this, simply supply the subject names with --s subject1 subject2 ... instead of --i. The segmentation will be stored as subject1/mri/sclimbic.mgz. This will automatically store a volume table (in FS stats format) in subject1/stats/sclimbic.stats; this stats file will automatically have the eTIV. After this, you can proceed with typical FreeSurfer ROI analysis, eg, a table with all the subjects can then be obtained with
asegstats2table --stats=sclimbic.stats -t sclimbic.dat --fsgd=group.fsgd
where it will get the list of subjects from the FSGD file. sclimbic.dat is the table of results. One can then do the group analysis with
mri_glmfit --table sclimbic.dat --fsgd group.fsgd --scale-by-etiv --o glm.results
Input with voxel sizes that are not 1mm Isotropic (--conform)
The network was trained on images with 1mm3 voxel size, so all inputs must conform to this constraint. If you input does not, you can add the --conform flag. This will resample the input to be 1mm3 and perform the segmentation in this space. When writing the output segmentation volume, the segmentation will be resampled back into the native resolution. This process may cause some degradation in the quality of the results.
Other options
--threads N : select the number of CPU threads to use
--cuda-device DeviceNo : use a GPU device. This does not speed things up all that much
--tal talextension : compute eTIV from an xfm file called input.talextension in the inputfolder --write_posteriors : write outposterior probability maps
Source Code
Source code is available at the FreeSurfer github page: https://github.com/freesurfer/freesurfer
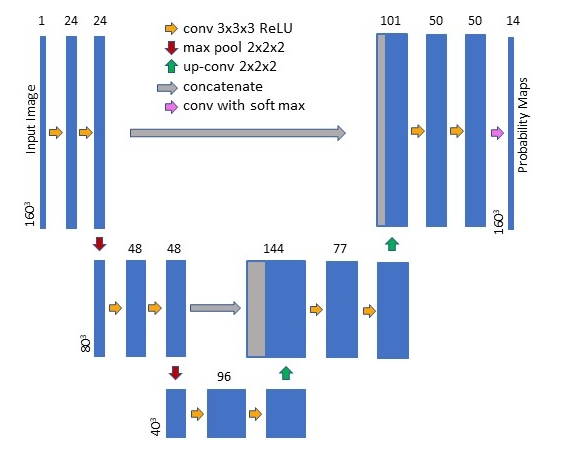
U-Net Training and Architecture
The U-Net was constructed and trained using the publicly available Neuron (Dalca, 2018) and Lab2Im (Billot et al., 2020a) python packages (both available at https://github.com/BBillot/hypothalamus_seg). The network was implemented in Keras (Chollet, 2016) with a Tensorflow (Abadi, 2016) backend. Our network architecture, augmentation, and training were identical to that of Billot et al., 2020a with the exception that we include intensity noise augmentation (i.e., the adding of white Gaussian noise to the image during training).