FreeSurfer For Martinos Users
Multiple FreeSurfer distributions can be accessed from any Linux computer on the Martinos network. This page will show you how to access a FreeSurfer distribution and point you towards some tutorials to help learn how to use it.
FreeSurfer Distributions
Important Note: When processing a group of subjects for your study, it is essential to process all your subjects with the same version of FreeSurfer, on the same OS platform and vendor, and for safety, even the same version of the OS. While we continue to work to ensure that results match across platforms, there are none-the-less system-level libraries that are OS dependent. An exception to this rule is that you may view and edit files across any platform or version, and run some post-processing tools (outside the recon-all stream) if you check with us first (for instance you may run the longitudinal processing with newer versions). You can assure that you are only using a particular version by copying $FREESURFER/build-stamp.txt to $SUBJECTS_DIR/fs-allowed-versions.txt
You may want to consult the freesurfer release notes to read about the changes in the current release(s) before choosing a version of Freesurfer to use FreeSurfer release notes. Should you wish to download and test a version of Freesurfer on your local machine, then please visit the download page FreeSurfer downloads
If you are about to process new data with FreeSurfer, then we recommend using the latest release. As of this writing, the latest Freesurfer release is version 8.1.0 (from July of 2025).
Official Freesurfer (stable) releases hosted internally: 8.1.0 (stable8.1.0) 8.0.0 (stable8.0.0) 7.4.1 (stable7.4.1) 7.4.0 (stable7.4.0) 7.3.2 (stable7.3.2) 7.2.0 (stable7.2.0) 7.1.1 (stable7.1.1) 6.0.1 6.0.0 (stable6) stable5_3_0 stable5_0_0 Non-release Freesurfer (unstable) development streams hosted internally: dev daily CentOS7 dev build for CentOS7, CentOS8, Rocky8 machines, MacOS 10.?? dev_cuda daily CentOS7 dev build for CentOS7, CentOS8, Rocky8 machines with cuda python packages dev_rocky8 daily Rocky8 dev build for Rocky8, Rocky9 machines (Freeview with ScribblePrompt) dev_rocky9 daily Rocky9 dev build for Rocky9 machines
Source FreeSurfer Environment
Choose a specific Freesurfer release from the list, use the following command to source your environment:
source /cluster/freesurfer/fsenv <release>
To use the 8.1.0 release as hosted internally on the Martinos Center servers, you would copy and paste this command into your terminal window:
Version 8.1.0 release:
source /cluster/freesurfer/fsenv 8.1.0
If you are in the process of analyzing data with the previous FreeSurfer version 6 release, then run this command in order to continue to work with version 6:
Version 6 release:
source /cluster/freesurfer/fsenv stable6
Development builds and releases on newer versions of Linux O/S's are built with newer versions of the g++/gcc compilers: gcc4 for CentOS7; gcc8 for CentOS8/Rocky8; gcc11 for Rocky9. In general, newer compiler versions can result in code that is better and more strictly compiled due to gcc toolchain bug fixes and adherence to the newer C/C++ standards. This can result in the compiled code being more optimized and efficient.
Development builds are a snapshot in time of ongoing development work that has not (yet) been fully tested like a stable numbered release. The contents of development builds change frequently (often on a daily basis) as part of accumulating the latest functionality pushed to the FreeSurfer Github repository. Therefore, PLEASE NOTE that development builds of Freesurfer are extremely unstable and should not be used for studies. You can browse the latest source code changes, enhancements and bug fixes via the FreeSurfer GitHub Actions tab.
Run the CentOS7 development build of freesurfer.
source /cluster/freesurfer/fsenv dev
Run the Rocky9 development build of freesurfer.
source /cluster/freesurfer/fsenv dev_rocky9
To ensure one of the above commands worked, type freeview into the command prompt, press enter, and see if FreeView opened. FreeView is a tool that comes with FreeSurfer - it lets you look at the data FreeSurfer creates along with many other files common to MRI research.
A Note On fs-*-env-autoselect Scripts
These scripts are the alternative ways to run Freesurfer releases 8.1.0, 8.0.0, 7.4.1, 7.1.1, 7.1.0, 6.0.0, and the non-release Freesurfer development stream.
source /usr/local/freesurfer/fs-stable810-env-autoselect (Release 8.1.0, stable8.1.0) source /usr/local/freesurfer/fs-stable800-env-autoselect (Release 8.0.0, stable8.0.0) source /usr/local/freesurfer/fs-stable741-env-autoselect (Release 7.4.1, stable7.4.1) source /usr/local/freesurfer/fs-stable711-env-autoselect (Release 7.1.1, stable7.1.1) source /usr/local/freesurfer/fs-stable710-env-autoselect (Release 7.1.0, stable7.1.0) source /usr/local/freesurfer/fs-stable6-env-autoselect (Release 6.0.0, stable6) source /usr/local/freesurfer/fs-dev-env-autoselect (Non-release development stream)
We have replaced them with a much simpler file structure. There will be no more additions to these scripts for any future or past releases.
A note on Bash vs TCSH
Most of FreeSurfer is used through typing commands into the terminal. Different labs use different terminal languages - these languages are called shells - some commands you type in to use FreeSurfer are slightly different depending on which shell your computer uses. Your computer probably uses Bash or TCSH, to figure that out enter the following command into the terminal.
echo $SHELL
The output will show a path, the final part of that path will be the name of your shell.
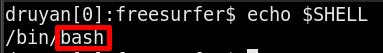
If you follow the FreeSurfer Tutorials to learn FreeSurfer, you will be guided through the differences in Bash and TCSH that matter to FreeSurfer. One example difference is how you would tell FreeSurfer where to save your processed subject data. Here is how that would look in each shell.
Bash
export SUBJECTS_DIR=/some/path_to/your/subjects
TCSH
setenv SUBJECTS_DIR /some/path_to/your/subjects
If you would like to change shell at any time, type in the name of the shell you would like into the terminal: bash or tcsh . Note that new terminal windows will still use your usual default shell.
Unpacking data from the scanners
When you finish scanning, you will "push" your data onto the Martinos DICOM server (bourget). "Unpacking" refers to the process of converting the data into a format that is useful for processing (such as NIFTI or MGZ). First, you have to find your data on the server. To do this, run
findsession subjectid
where "subjectid" is the code that you used when you registered the subject on the scanner. You can also use any subset of characters. There are other options that you can find by running findsession with -h. When you run findsession and locate your subject, you will see several fields. Locate the one called "PATH". Eg, it may look something like /cluster/archive/296/siemens/Avanto-25096-20111215-151633-370000. Once you have this, you can run something like
dcmunpack -src /cluster/archive/296/siemens/Avanto-25096-20111215-151633-370000 -trg /path/to/your/output -auto-runseq mgz -no-exit-on-error
This will convert all the scans to mgz (ie, FreeSurfer) format and store them in /path/to/your/output. There are other options for dcmunpack which can be found by running it with -help. If you are using FreeSurfer 7.4.X, then dcmunpack will automatically convert DICOMs created on the new (as of 10/23) XA scanners. If you are using 7.3.X, then you will need to add -dcm2niix to the command line. Previous versions of FreeSurfer will not be able to unpack the XA data.
Learning how to use FreeSurfer
Here is a recommended way to get up to speed on using FreeSurfer:
If you are not familiar with using a Unix terminal, watch this video: https://www.youtube.com/watch?v=6eJMxh7PlOY
Here is a link to online versions of a FreeSurfer course: https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial
Each lecture has a corresponding video found here: https://www.youtube.com/channel/UCruQerP8aa-gYttXkAcyveA
To get a basic understanding of FreeSurfer it is recommended to go through all course material up to and including "Quality Checking a Recon". From there you can complete the rest to explore different specific functions of FreeSurfer, or select tutorials related to the research you plan to do.
Processing your first subject
If you have data you would like to process with FreeSurfer, you can do so using the command recon-all. An example of how to run this command is below:
First, make sure your computer is set up to run FreeSurfer. Follow the accessing FreeSurfer section above to do so.
Choose a directory where you would like FreeSurfer to store your processed data. See the A note on Bash vs TCSH section above to do so.
Navigate to the directory of your DICOM image data with cd /path/to/data/here , then run the following command:
recon-all -all -i <one slice in the anatomical dicom series> -s <subject id that you make up>
Things to note:
-all will run all the steps in the FreeSurfer processing stream. Alternatively, you can run different parts of the stream.
-i stands for input. Here, you would specify one dicom file/slice in the MPRAGE scan series you collected ( FreeSurfer will find the rest of the slices automatically). You could also specify nifti files as input. If you have more than one MPRAGE or input for a given subject, use another -i flag for each one.
-s specifies the name of the subject and will create a directory with that name for all the subject's FreeSurfer output.
The process is complete when the last line printed to the screen (and in the recon-all.log found in the subject's scripts directory) says "recon-all exited without errors". If you run into errors, search the mailing list for help or follow the steps in BugReporting.
General Linux Help
Examples of scripts and aliases that can make it easier to source, use, and process with FreeSurfer can be found here.
Some basic information on text editors can be found here.
Need help?
There is a wealth of knowledge here on the FSWiki. On the top right hand corner is a search bar - be sure to press the Text option when searching, it offers more results than Topic.
Other resources include an online support archive, see here for more details: https://surfer.nmr.mgh.harvard.edu/fswiki/FreeSurferSupport
Other resources
You may find these wiki pages helpful as you get started:
